Difference between revisions of "AddA"
| Line 54: | Line 54: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | * ''[[recO]] [[addA]]-[[addB]]'' double mutants are extremely sensitive against DNA damaging agents {{PubMed|26001966}} | ||
=== Database entries === | === Database entries === | ||
| Line 147: | Line 148: | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>,8387145,15610857,7746142, 19129187 15009890 10756102 9781875 16632468 17570399 20350930 21809208 21821766 22307084 24682829 24670664 22383849 21071401 8752329 8752323 25939832</pubmed> | + | <pubmed>,8387145,15610857,7746142, 19129187 15009890 10756102 9781875 16632468 17570399 20350930 21809208 21821766 22307084 24682829 24670664 22383849 21071401 8752329 8752323 25939832 26001966</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 08:04, 7 July 2015
- Description: ATP-dependent deoxyribonuclease (subunit A), required for efficient survival and replication restart after replication-transcription conflicts
| Gene name | addA |
| Synonyms | recE5 |
| Essential | no |
| Product | ATP-dependent deoxyribonuclease (subunit A)) |
| Function | DNA repair/ recombination |
| Gene expression levels in SubtiExpress: addA | |
| Interactions involving this protein in SubtInteract: AddA | |
| MW, pI | 140 kDa, 5.127 |
| Gene length, protein length | 3696 bp, 1232 aa |
| Immediate neighbours | addB, sbcD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 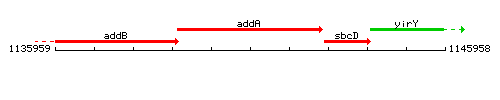
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed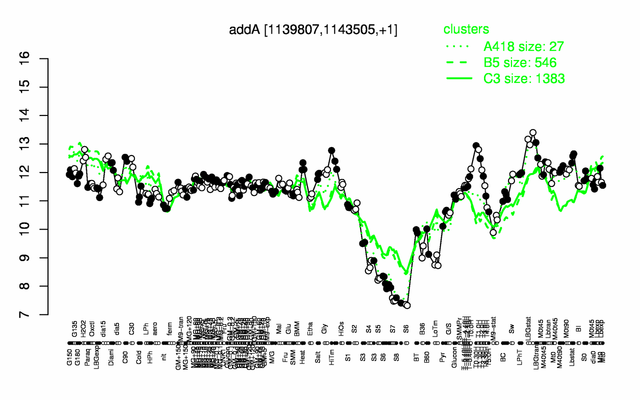
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, genetic competence
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU10630
Phenotypes of a mutant
Database entries
- BsubCyc: BSU10630
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- the enzyme is functional as a heterodimer of the AddA and AddB subunits, that it is a rapid and processive DNA helicase, and that it catalyses DNA unwinding using one single-stranded DNA motor of 3'→5' polarity located in the AddA subunit PubMed
- the AddB subunit contains a second putative ATP-binding pocket, but this does not contribute to the observed helicase activity and may instead be involved in the recognition of recombination hotspot sequences PubMed
- Protein family: uvrD-like helicase C-terminal domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU10630
- UniProt: P23478
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 147 PubMed
Biological materials
- Mutant: GP1106 (addA-addB, spc), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Mark Dillingham, Bristol, U.K. (homepage)
Your additional remarks
References
Reviews
Original publications
Begoña Carrasco, Tribhuwan Yadav, Ester Serrano, Juan C Alonso
Bacillus subtilis RecO and SsbA are crucial for RecA-mediated recombinational DNA repair.
Nucleic Acids Res: 2015, 43(12);5984-97
[PubMed:26001966]
[WorldCat.org]
[DOI]
(I p)
Samuel Million-Weaver, Ariana Nakta Samadpour, Houra Merrikh
Replication Restart after Replication-Transcription Conflicts Requires RecA in Bacillus subtilis.
J Bacteriol: 2015, 197(14);2374-82
[PubMed:25939832]
[WorldCat.org]
[DOI]
(I p)
Neville S Gilhooly, Mark S Dillingham
Recombination hotspots attenuate the coupled ATPase and translocase activities of an AddAB-type helicase-nuclease.
Nucleic Acids Res: 2014, 42(9);5633-43
[PubMed:24682829]
[WorldCat.org]
[DOI]
(I p)
Wojciech W Krajewski, Xin Fu, Martin Wilkinson, Nora B Cronin, Mark S Dillingham, Dale B Wigley
Structural basis for translocation by AddAB helicase-nuclease and its arrest at χ sites.
Nature: 2014, 508(7496);416-9
[PubMed:24670664]
[WorldCat.org]
[DOI]
(I p)
Pierre Nicolas, Ulrike Mäder, Etienne Dervyn, Tatiana Rochat, Aurélie Leduc, Nathalie Pigeonneau, Elena Bidnenko, Elodie Marchadier, Mark Hoebeke, Stéphane Aymerich, Dörte Becher, Paola Bisicchia, Eric Botella, Olivier Delumeau, Geoff Doherty, Emma L Denham, Mark J Fogg, Vincent Fromion, Anne Goelzer, Annette Hansen, Elisabeth Härtig, Colin R Harwood, Georg Homuth, Hanne Jarmer, Matthieu Jules, Edda Klipp, Ludovic Le Chat, François Lecointe, Peter Lewis, Wolfram Liebermeister, Anika March, Ruben A T Mars, Priyanka Nannapaneni, David Noone, Susanne Pohl, Bernd Rinn, Frank Rügheimer, Praveen K Sappa, Franck Samson, Marc Schaffer, Benno Schwikowski, Leif Steil, Jörg Stülke, Thomas Wiegert, Kevin M Devine, Anthony J Wilkinson, Jan Maarten van Dijl, Michael Hecker, Uwe Völker, Philippe Bessières, Philippe Noirot
Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis.
Science: 2012, 335(6072);1103-6
[PubMed:22383849]
[WorldCat.org]
[DOI]
(I p)
Kayarat Saikrishnan, Joseph T Yeeles, Neville S Gilhooly, Wojciech W Krajewski, Mark S Dillingham, Dale B Wigley
Insights into Chi recognition from the structure of an AddAB-type helicase-nuclease complex.
EMBO J: 2012, 31(6);1568-78
[PubMed:22307084]
[WorldCat.org]
[DOI]
(I p)
Christopher T Brown, Laura K Fishwick, Binna M Chokshi, Marissa A Cuff, Jay M Jackson, Travis Oglesby, Alison T Rioux, Enrique Rodriguez, Gregory S Stupp, Austin H Trupp, James S Woollcombe-Clarke, Tracy N Wright, William J Zaragoza, Jennifer C Drew, Eric W Triplett, Wayne L Nicholson
Whole-genome sequencing and phenotypic analysis of Bacillus subtilis mutants following evolution under conditions of relaxed selection for sporulation.
Appl Environ Microbiol: 2011, 77(19);6867-77
[PubMed:21821766]
[WorldCat.org]
[DOI]
(I p)
Natalia Fili, Christopher P Toseland, Mark S Dillingham, Martin R Webb, Justin E Molloy
A single-molecule approach to visualize the unwinding activity of DNA helicases.
Methods Mol Biol: 2011, 778;193-214
[PubMed:21809208]
[WorldCat.org]
[DOI]
(I p)
Joseph T P Yeeles, Emma J Gwynn, Martin R Webb, Mark S Dillingham
The AddAB helicase-nuclease catalyses rapid and processive DNA unwinding using a single Superfamily 1A motor domain.
Nucleic Acids Res: 2011, 39(6);2271-85
[PubMed:21071401]
[WorldCat.org]
[DOI]
(I p)
Natali Fili, Gregory I Mashanov, Christopher P Toseland, Christopher Batters, Mark I Wallace, Joseph T P Yeeles, Mark S Dillingham, Martin R Webb, Justin E Molloy
Visualizing helicases unwinding DNA at the single molecule level.
Nucleic Acids Res: 2010, 38(13);4448-57
[PubMed:20350930]
[WorldCat.org]
[DOI]
(I p)
Joseph T P Yeeles, Richard Cammack, Mark S Dillingham
An iron-sulfur cluster is essential for the binding of broken DNA by AddAB-type helicase-nucleases.
J Biol Chem: 2009, 284(12);7746-55
[PubMed:19129187]
[WorldCat.org]
[DOI]
(P p)
Joseph T P Yeeles, Mark S Dillingham
A dual-nuclease mechanism for DNA break processing by AddAB-type helicase-nucleases.
J Mol Biol: 2007, 371(1);66-78
[PubMed:17570399]
[WorldCat.org]
[DOI]
(P p)
Frédéric Chédin, Naofumi Handa, Mark S Dillingham, Stephen C Kowalczykowski
The AddAB helicase/nuclease forms a stable complex with its cognate chi sequence during translocation.
J Biol Chem: 2006, 281(27);18610-7
[PubMed:16632468]
[WorldCat.org]
[DOI]
(P p)
Alexander Serganov, Yu-Ren Yuan, Olga Pikovskaya, Anna Polonskaia, Lucy Malinina, Anh Tuân Phan, Claudia Hobartner, Ronald Micura, Ronald R Breaker, Dinshaw J Patel
Structural basis for discriminative regulation of gene expression by adenine- and guanine-sensing mRNAs.
Chem Biol: 2004, 11(12);1729-41
[PubMed:15610857]
[WorldCat.org]
[DOI]
(P p)
Etienne Dervyn, Marie-Françoise Noirot-Gros, Peggy Mervelet, Steven McGovern, S Dusko Ehrlich, Patrice Polard, Philippe Noirot
The bacterial condensin/cohesin-like protein complex acts in DNA repair and regulation of gene expression.
Mol Microbiol: 2004, 51(6);1629-40
[PubMed:15009890]
[WorldCat.org]
[DOI]
(P p)
F Chédin, S D Ehrlich, S C Kowalczykowski
The Bacillus subtilis AddAB helicase/nuclease is regulated by its cognate Chi sequence in vitro.
J Mol Biol: 2000, 298(1);7-20
[PubMed:10756102]
[WorldCat.org]
[DOI]
(P p)
F Chédin, P Noirot, V Biaudet, S D Ehrlich
A five-nucleotide sequence protects DNA from exonucleolytic degradation by AddAB, the RecBCD analogue of Bacillus subtilis.
Mol Microbiol: 1998, 29(6);1369-77
[PubMed:9781875]
[WorldCat.org]
[DOI]
(P p)
B J Haijema, R Meima, J Kooistra, G Venema
Effects of lysine-to-glycine mutations in the ATP-binding consensus sequences in the AddA and AddB subunits on the Bacillus subtilis AddAB enzyme activities.
J Bacteriol: 1996, 178(17);5130-7
[PubMed:8752329]
[WorldCat.org]
[DOI]
(P p)
B J Haijema, G Venema, J Kooistra
The C terminus of the AddA subunit of the Bacillus subtilis ATP-dependent DNase is required for the ATP-dependent exonuclease activity but not for the helicase activity.
J Bacteriol: 1996, 178(17);5086-91
[PubMed:8752323]
[WorldCat.org]
[DOI]
(P p)
B J Haijema, L W Hamoen, J Kooistra, G Venema, D van Sinderen
Expression of the ATP-dependent deoxyribonuclease of Bacillus subtilis is under competence-mediated control.
Mol Microbiol: 1995, 15(2);203-11
[PubMed:7746142]
[WorldCat.org]
[DOI]
(P p)
J Kooistra, B J Haijema, G Venema
The Bacillus subtilis addAB genes are fully functional in Escherichia coli.
Mol Microbiol: 1993, 7(6);915-23
[PubMed:8387145]
[WorldCat.org]
[DOI]
(P p)