Difference between revisions of "EpsI"
(→Expression and regulation) |
|||
| Line 113: | Line 113: | ||
** induction by sequestration of [[SinR]] by [[SinI]] or [[SlrA]] {{PubMed|15661000,19788541}} | ** induction by sequestration of [[SinR]] by [[SinI]] or [[SlrA]] {{PubMed|15661000,19788541}} | ||
** the ''[[epsA]]-[[epsB]]-[[epsC]]-[[epsD]]-[[epsE]]-[[epsF]]-[[epsG]]-[[epsH]]-[[epsI]]-[[epsJ]]-[[epsK]]-[[epsL]]-[[epsM]]-[[epsN]]-[[epsO]]'' operon is not expressed in a ''[[ymdB]]'' mutant {{PubMed|21856853}} | ** the ''[[epsA]]-[[epsB]]-[[epsC]]-[[epsD]]-[[epsE]]-[[epsF]]-[[epsG]]-[[epsH]]-[[epsI]]-[[epsJ]]-[[epsK]]-[[epsL]]-[[epsM]]-[[epsN]]-[[epsO]]'' operon is not expressed in a ''[[ymdB]]'' mutant {{PubMed|21856853}} | ||
| − | ** the amount of the mRNA is substantially decreased upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | + | ** the amount of the mRNA is substantially decreased upon depletion of [[Rny|RNase Y]] (this is likely due to the increased stability of the ''[[sinR]]'' mRNA) {{PubMed|21815947}} |
| − | ** the [[EAR riboswitch]] (eps-associated [[RNA switch]]) located between'' [[epsB]]'' and ''[[epsC]]'' mediates processive antitermination and allows expression of the long eps operon {{PubMed|20374491}} | + | ** the [[EAR riboswitch]] (eps-associated [[RNA switch]]) located between'' [[epsB]]'' and ''[[epsC]]'' mediates processive antitermination and allows expression of the long eps operon {{PubMed|20374491}} |
=Biological materials = | =Biological materials = | ||
Revision as of 14:06, 20 November 2011
- Description: extracellular polysaccharide synthesis
| Gene name | epsI |
| Synonyms | yveS |
| Essential | no |
| Product | unknown |
| Function | biofilm formation |
| Regulation of this protein in SubtiPathways: Biofilm | |
| MW, pI | 41 kDa, 8.043 |
| Gene length, protein length | 1074 bp, 358 aa |
| Immediate neighbours | epsJ, epsH |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 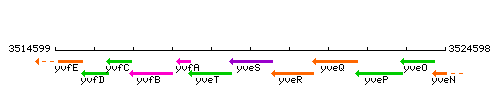
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
AbrB regulon, EAR riboswitch, SinR regulon
The gene
Basic information
- Locus tag: BSU34290
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: polysaccharide pyruvyl transferase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P71058
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
- induction by sequestration of SinR by SinI or SlrA PubMed
- the epsA-epsB-epsC-epsD-epsE-epsF-epsG-epsH-epsI-epsJ-epsK-epsL-epsM-epsN-epsO operon is not expressed in a ymdB mutant PubMed
- the amount of the mRNA is substantially decreased upon depletion of RNase Y (this is likely due to the increased stability of the sinR mRNA) PubMed
- the EAR riboswitch (eps-associated RNA switch) located between epsB and epsC mediates processive antitermination and allows expression of the long eps operon PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Richard Losick, Harvard Univ., Cambridge, USA homepage
Your additional remarks
References
Reviews
Original publications
The EAR RNA switch
Irnov Irnov, Wade C Winkler
A regulatory RNA required for antitermination of biofilm and capsular polysaccharide operons in Bacillales.
Mol Microbiol: 2010, 76(3);559-75
[PubMed:20374491]
[WorldCat.org]
[DOI]
(I p)
Zasha Weinberg, Joy X Wang, Jarrod Bogue, Jingying Yang, Keith Corbino, Ryan H Moy, Ronald R Breaker
Comparative genomics reveals 104 candidate structured RNAs from bacteria, archaea, and their metagenomes.
Genome Biol: 2010, 11(3);R31
[PubMed:20230605]
[WorldCat.org]
[DOI]
(I p)
Other original publications
Additional publications: PubMed
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Diethmaier C, Pietack N, Gunka K, Wrede C, Lehnik-Habrink M, Herzberg C, Hübner S, Stülke J A Novel Factor Controlling Bistability in Bacillus subtilis: The YmdB Protein Affects Flagellin Expression and Biofilm Formation. J Bacteriol.: 2011, 193(21):5997-6007. PubMed:21856853