Difference between revisions of "AddA"
(→Labs working on this gene/protein) |
|||
| Line 57: | Line 57: | ||
=== Additional information=== | === Additional information=== | ||
| − | + | * A mutation was found in this gene after evolution under relaxed selection for sporulation {{PubMed|21821766}} | |
| − | |||
| Line 139: | Line 138: | ||
==Original publications== | ==Original publications== | ||
'''Additional publications:''' {{PubMed|21071401}} | '''Additional publications:''' {{PubMed|21071401}} | ||
| − | <pubmed>,8387145,15610857,7746142, 19129187 15009890 10756102 9781875 16632468 17570399 20350930 21809208</pubmed> | + | <pubmed>,8387145,15610857,7746142, 19129187 15009890 10756102 9781875 16632468 17570399 20350930 21809208 21821766</pubmed> |
'''Additional publications:''' {{PubMed|21071401}} | '''Additional publications:''' {{PubMed|21071401}} | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:52, 15 August 2011
- Description: ATP-dependent deoxyribonuclease (subunit A)
| Gene name | addA |
| Synonyms | recE5 |
| Essential | no |
| Product | ATP-dependent deoxyribonuclease (subunit A)) |
| Function | DNA repair/ recombination |
| Interactions involving this protein in SubtInteract: AddA | |
| MW, pI | 140 kDa, 5.127 |
| Gene length, protein length | 3696 bp, 1232 aa |
| Immediate neighbours | addB, sbcD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 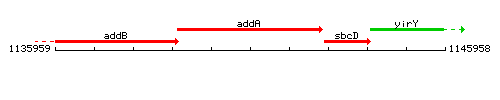
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, genetic competence
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU10630
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- the enzyme is functional as a heterodimer of the AddA and AddB subunits, that it is a rapid and processive DNA helicase, and that it catalyses DNA unwinding using one single-stranded DNA motor of 3'→5' polarity located in the AddA subunit PubMed
- the AddB subunit contains a second putative ATP-binding pocket, but this does not contribute to the observed helicase activity and may instead be involved in the recognition of recombination hotspot sequences PubMed
- Protein family: uvrD-like helicase C-terminal domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P23478
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP1106 (addAB, spc), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Mark Dillingham, Bristol, U.K. (homepage)
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed
Additional publications: PubMed