Difference between revisions of "PdxS"
| Line 93: | Line 93: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
| − | ** [[PdxS]]-[[PdxT]] | + | ** [[PdxS]]-[[PdxT]] {{PubMed|25473090,14762015,16030023,17159152}} |
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
| Line 153: | Line 153: | ||
=References= | =References= | ||
| − | <pubmed>15911615,14762015,,14651647, 17726680, 16493705, 16030023, 19152323,17159152 22517742 15378759</pubmed> | + | <pubmed>15911615,14762015,25473090,14651647, 17726680, 16493705, 16030023, 19152323,17159152 22517742 15378759</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 09:26, 5 December 2014
- Description: pyridoxal-5'-phosphate synthase (synthase domain)
| Gene name | pdxS |
| Synonyms | yaaD |
| Essential | no |
| Product | pyridoxal-5'-phosphate synthase (synthase domain) |
| Function | pyridoxal-5'-phosphate biosynthesis |
| Gene expression levels in SubtiExpress: pdxS | |
| Interactions involving this protein in SubtInteract: PdxS | |
| Metabolic function and regulation of this protein in SubtiPathways: PdxS | |
| MW, pI | 31 kDa, 5.085 |
| Gene length, protein length | 882 bp, 294 aa |
| Immediate neighbours | dacA, pdxT |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 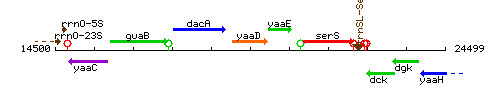
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed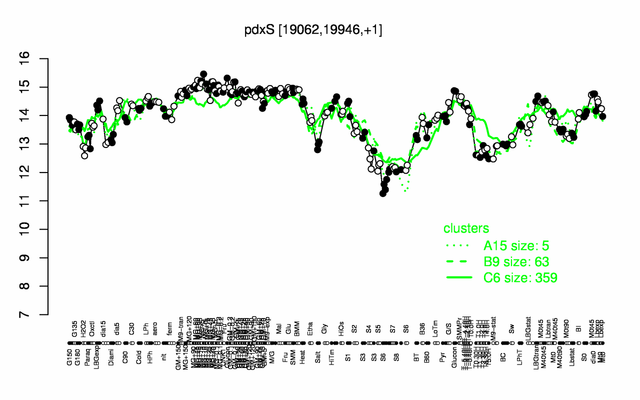
| |
Contents
Categories containing this gene/protein
biosynthesis of cofactors, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU00110
Phenotypes of a mutant
auxotrophic for pyridoxal 5'-phosphate PubMed
Database entries
- BsubCyc: BSU00110
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: pdxS/SNZ family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Effectors of protein activity:
Database entries
- BsubCyc: BSU00110
- UniProt: P37527
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 6001 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 16290 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 7634 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 4624 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 7523 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References