Difference between revisions of "ClpC"
| Line 88: | Line 88: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' AAA-ATPase [http://pfam.sanger.ac.uk/family?acc=PF07724 PFAM] | + | * '''[[Domains]]:''' AAA-ATPase [http://pfam.sanger.ac.uk/family?acc=PF07724 PFAM] |
* '''Modification:''' | * '''Modification:''' | ||
** phosphorylated on Arg-5 and Arg-254 {{PubMed|22517742}} | ** phosphorylated on Arg-5 and Arg-254 {{PubMed|22517742}} | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 166: | Line 166: | ||
<pubmed> 17302811 23375660 23479438,19609260,19781636</pubmed> | <pubmed> 17302811 23375660 23479438,19609260,19781636</pubmed> | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>9987115,8016067,9000055,12923101,10447896,9141693,2113920,16497325,19226326,8793870,10809708,14679237,17560370,11684022,8195092,11722737,11914365,12028382,18689476,19361434,9890793, 19767395 ,9987115, 11544224, 17981983, 14763982, 8016066 19361434 18689473 20070525 20923420 20852588 22517742 21622759 21368759 21821766 23595989 18786145,16525504,17380125, 16163393,12598648</pubmed> | + | <pubmed>9987115,8016067,9000055,12923101,10447896,9141693,2113920,16497325,19226326,8793870,10809708,14679237,17560370,11684022,8195092,11722737,11914365,12028382,18689476,19361434,9890793, 19767395 ,9987115, 11544224, 17981983, 14763982, 8016066 19361434 18689473 20070525 20923420 20852588 22517742 21622759 21368759 21821766 23595989 18786145,16525504,17380125, 16163393,12598648 24263382</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 07:30, 16 December 2013
- Description: ATPase subunit of the ATP-dependent ClpC-ClpP protease, involved in competence development, heat shock regulation, motility, sporulation, protein quality control, biofilm formation
| Gene name | clpC |
| Synonyms | mecB |
| Essential | no |
| Product | ATPase subunit of the ClpC-ClpP protease |
| Function | protein degradation positive regulator of autolysin (LytC and LytD) synthesis |
| Gene expression levels in SubtiExpress: clpC | |
| Interactions involving this protein in SubtInteract: ClpC | |
| Metabolic function and regulation of this protein in SubtiPathways: Stress | |
| MW, pI | 89 kDa, 5.746 |
| Gene length, protein length | 2430 bp, 810 aa |
| Immediate neighbours | mcsB, radA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 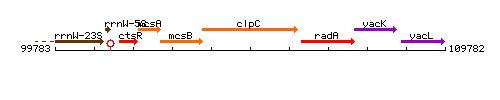
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed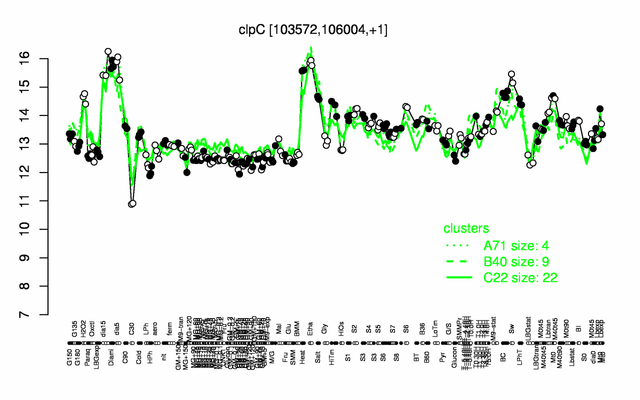
| |
Contents
Categories containing this gene/protein
proteolysis, sporulation proteins, general stress proteins (controlled by SigB), heat shock proteins, phosphoproteins
This gene is a member of the following regulons
CtsR regulon, SigB regulon, SigF regulon
The gene
Basic information
- Locus tag: BSU00860
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATPase/chaperone
- Protein family: mecA family (according to Swiss-Prot) clpA/clpB family. ClpC subfamily (according to Swiss-Prot), AAA+ -type ATPase (IPR013093) InterPro (PF07724) PFAM
Targets of ClpC-ClpP-dependent protein degradation
Extended information on the protein
- Kinetic information:
- Modification:
- phosphorylated on Arg-5 and Arg-254 PubMed
- Effectors of protein activity:
Database entries
- UniProt: P37571
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Expression vector:
- lacZ fusion:
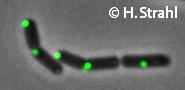
- GFP fusion: C-terminal GFP fusions (single copy, also as CFP and YFP variants) available from the Hamoen Lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Newcastle University, UK homepage
Kürsad Turgay, Freie Universität Berlin, Germany homepage
Your additional remarks
References
Reviews
Original Publications