Difference between revisions of "PpsC"
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || production of the antibacterial compound plipastatin | |style="background:#ABCDEF;" align="center"|'''Function''' || production of the antibacterial compound plipastatin | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://cellpublisher.gobics.de/subtiexpress/ ''Subti''Express]''': [http://cellpublisher.gobics.de/subtiexpress/bsu/BSU18320 ppsC] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 287 kDa, 5.352 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 287 kDa, 5.352 | ||
Revision as of 10:16, 7 August 2012
- Description: plipastatin synthetase
| Gene name | ppsC |
| Synonyms | |
| Essential | no |
| Product | plipastatin synthetase |
| Function | production of the antibacterial compound plipastatin |
| Gene expression levels in SubtiExpress: ppsC | |
| MW, pI | 287 kDa, 5.352 |
| Gene length, protein length | 7665 bp, 2555 aa |
| Immediate neighbours | ppsD, ppsB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 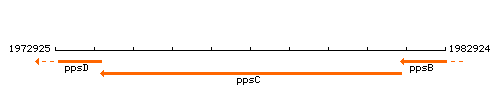
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed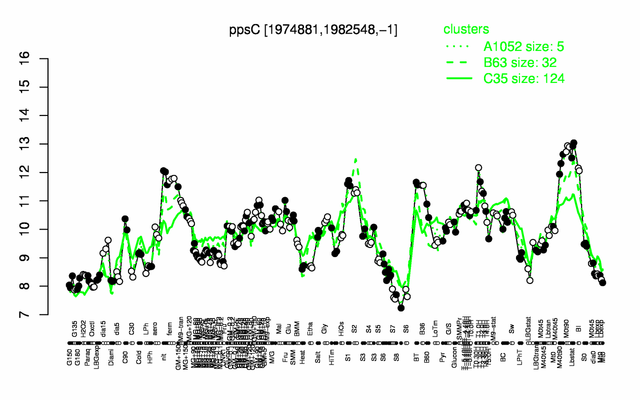
| |
Contents
Categories containing this gene/protein
miscellaneous metabolic pathways, biosynthesis of antibacterial compounds
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18320
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: ATP-dependent AMP-binding enzyme family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- PpsC is a member of a suspected group of hubs proteins that were suggested to be involved in a large number of interactions PubMed
Database entries
- Structure:
- UniProt: P39847
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Elodie Marchadier, Rut Carballido-López, Sophie Brinster, Céline Fabret, Peggy Mervelet, Philippe Bessières, Marie-Françoise Noirot-Gros, Vincent Fromion, Philippe Noirot
An expanded protein-protein interaction network in Bacillus subtilis reveals a group of hubs: Exploration by an integrative approach.
Proteomics: 2011, 11(15);2981-91
[PubMed:21630458]
[WorldCat.org]
[DOI]
(I p)
K Tsuge, T Ano, M Hirai, Y Nakamura, M Shoda
The genes degQ, pps, and lpa-8 (sfp) are responsible for conversion of Bacillus subtilis 168 to plipastatin production.
Antimicrob Agents Chemother: 1999, 43(9);2183-92
[PubMed:10471562]
[WorldCat.org]
[DOI]
(P p)
Valentina Tosato, Alessandra M Albertini, Michela Zotti, Sabrina Sonda, Carlo V Bruschi
Sequence completion, identification and definition of the fengycin operon in Bacillus subtilis 168.
Microbiology (Reading): 1997, 143 ( Pt 11);3443-3450
[PubMed:9387222]
[WorldCat.org]
[DOI]
(P p)