Difference between revisions of "GudB"
(→References) |
|||
| Line 143: | Line 143: | ||
=References= | =References= | ||
| + | ==Reviews== | ||
| + | <pubmed>19698086 8299344 7705101 19895831</pubmed> | ||
| + | ==Original publications== | ||
'''Additional publications:''' {{PubMed|22178969}} | '''Additional publications:''' {{PubMed|22178969}} | ||
<pubmed>18603778,9829940 ,17183217 18723616, 18326565 20630473 17981983 21219666 22178973 </pubmed> | <pubmed>18603778,9829940 ,17183217 18723616, 18326565 20630473 17981983 21219666 22178973 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:13, 26 December 2011
- Description: trigger enzyme: glutamate dehydrogenase (cryptic in 168 and derivatives)
| Gene name | gudB |
| Synonyms | ypcA |
| Essential | no |
| Product | trigger enzyme: glutamate dehydrogenase |
| Function | glutamate utilization, control of GltC activity |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 47 kDa, 5.582 |
| Gene length, protein length | 1278 bp, 426 aa |
| Immediate neighbours | ypdA, ypbH |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 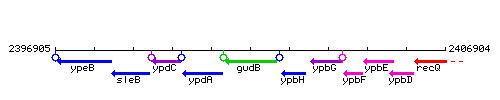
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of amino acids, transcription factors and their control, trigger enzyme
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22960
Phenotypes of a mutant
- The gene is cryptic. If gudB is activated (gudB1 mutation), the bacteria are able to utilize glutamate as the only carbon source. PubMed
- transcription profile of a rocG gudB mutant strain: GEO PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-glutamate + H2O + NAD+ = 2-oxoglutarate + NH3 + NADH (according to Swiss-Prot)
- Protein family: Glu/Leu/Phe/Val dehydrogenases family (according to Swiss-Prot)
- Paralogous protein(s): RocG
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- UniProt: P50735
- KEGG entry: [4]
- E.C. number: 1.4.1.2
Additional information
Expression and regulation
- Operon: gudB PubMed
- Regulation: constitutively expressed
- Regulatory mechanism:
- Additional information: GudB is subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant: GP691 (cat), GP1160 (del aphA3) both available in Stülke lab
- Expression vector:
- for purification of GudB from E. coli carrying an N-terminal Strep-tag: pGP863 (in pGP172) available in Stülke lab
- for purification of GudB1 from E. coli carrying an N-terminal Strep-tag: pGP864 (in pGP172) available in Stülke lab
- for ectopic expression of gudB with its native promoter: pGP900 (in pAC5), available in Stülke lab
- wild type gudB, expression in B. subtilis, in pBQ200: pGP1712, available in Stülke lab
- lacZ fusion: pGP651 (in pAC5), available in Stülke lab
- FLAG-tag construct: GP1194 (gudB, spc, based on pGP1331), GP1195 (gudB1, spc, based on pGP1331), available in Stülke lab
- GFP fusion:
- two-hybrid system:
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
The GudB protein is active in other legacy B. subtilis strains (e.g. strain 122). Thus, it can be speculated that the ancestral gudB gene was not cryptic, but became so as a product of the "domestication" of B. subtilis 168 in the lab. PubMed
References
Reviews
Jason R Treberg, Margaret E Brosnan, Malcolm Watford, John T Brosnan
On the reversibility of glutamate dehydrogenase and the source of hyperammonemia in the hyperinsulinism/hyperammonemia syndrome.
Adv Enzyme Regul: 2010, 50(1);34-43
[PubMed:19895831]
[WorldCat.org]
[DOI]
(I p)
Victoria I Bunik, Alisdair R Fernie
Metabolic control exerted by the 2-oxoglutarate dehydrogenase reaction: a cross-kingdom comparison of the crossroad between energy production and nitrogen assimilation.
Biochem J: 2009, 422(3);405-21
[PubMed:19698086]
[WorldCat.org]
[DOI]
(I e)
N M Brunhuber, J S Blanchard
The biochemistry and enzymology of amino acid dehydrogenases.
Crit Rev Biochem Mol Biol: 1994, 29(6);415-67
[PubMed:7705101]
[WorldCat.org]
[DOI]
(P p)
R C Hudson, R M Daniel
L-glutamate dehydrogenases: distribution, properties and mechanism.
Comp Biochem Physiol B: 1993, 106(4);767-92
[PubMed:8299344]
[WorldCat.org]
[DOI]
(P p)
Original publications
Additional publications: PubMed