Difference between revisions of "PyrR"
| Line 80: | Line 80: | ||
* '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P39765 P39765] | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P39765 P39765] | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU15470] | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU15470 BSU15470] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/2.4.2.9 2.4.2.9] | * '''E.C. number:''' [http://www.expasy.org/enzyme/2.4.2.9 2.4.2.9] | ||
Revision as of 23:47, 13 May 2009
- Description: transcriptional antiterminator of the pyr operon
| Gene name | pyrR |
| Synonyms | |
| Essential | no |
| Product | transcriptional antiterminator with minor uracil phosphoribosyltransferase activity |
| Function | regulation of pyrimidine biosynthesis |
| MW, pI | 20 kDa, 4.991 |
| Gene length, protein length | 543 bp, 181 aa |
| Immediate neighbours | ylyB, pyrP |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 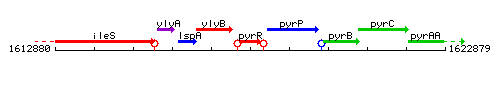
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: PyrR subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Structure: 1A3C (dimeric form)
- Swiss prot entry: P39765
- KEGG entry: BSU15470
- E.C. number: 2.4.2.9
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Tomchik, D. R., Turner, R. J., Switzer, R. L., and Smith, J. L. (1998) Adaptation of an enzyme to regulatory function: structure of Bacillus subtilis PyrR, a pyr RNA-binding attenuation protein and uracil phosphoribosyltransferase. Structure 6: 337-350. PubMed
- Turner, R. J., Bonner, E. R., Grabner, G. K., and Switzer, R. L. (1998) Purification and characterization of Bacillus subtilis PyrR, a bifunctional pyr mRNA-binding attenuation protein/uracil phosphoribosyltransferase. J Biol Chem 273: 5932-5938. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed