Difference between revisions of "YhaM"
| Line 1: | Line 1: | ||
| − | * '''Description:''' RNase, | + | * '''Description:''' RNase, 3'-> 5' exoribonuclease <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 115: | Line 115: | ||
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| + | |||
| + | [[David Bechhofer]], Mount Sinai School, New York, USA [http://www.mountsinai.org/Research/Centers%20Laboratories%20and%20Programs/Bechhofer%20Laboratory?citype=Physician&ciid=Bechhofer%20David%20H%201255565 Homepage] | ||
=Your additional remarks= | =Your additional remarks= | ||
Revision as of 20:36, 8 April 2009
- Description: RNase, 3'-> 5' exoribonuclease
| Gene name | yhaM |
| Synonyms | |
| Essential | no |
| Product | Rnase YhaM |
| Function | RNA degradation |
| MW, pI | 35 kDa, 5.851 |
| Gene length, protein length | 942 bp, 314 aa |
| Immediate neighbours | yhaN, yhaL |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 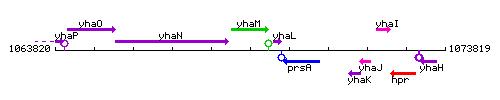
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 3'-5' exoribonuclease
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: Cytoplasm (Homogeneous) PubMed
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
David Bechhofer, Mount Sinai School, New York, USA Homepage
Your additional remarks
References
- Gerth et al. (2008) Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis. J Bacteriol 190:321-331 PubMed
- Meile et al. (2006) Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory Proteomics 6: 2135-2146. PubMed
- Oussenko, I. A., Abe, T., Ujiie, H., Muto, A. & Bechhofer, D. H. (2005). Participation of 3'-to-5' exoribonucleases in the turnover of Bacillus subtilis mRNA. J Bacteriol 187:2758-2767. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed