Difference between revisions of "IolD"
| Line 93: | Line 93: | ||
* '''Sigma factor:''' [[SigA]] | * '''Sigma factor:''' [[SigA]] | ||
| − | * '''Regulation:''' repressed by glucose ([[CcpA]]) | + | * '''Regulation:''' repressed by glucose ([[CcpA]]) |
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| Line 119: | Line 119: | ||
=References= | =References= | ||
| − | # Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in ''Bacillus subtilis'': regulation of the central metabolic pathways. ''Metab Eng.'' '''5:''' 133-149 | + | # Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in ''Bacillus subtilis'': regulation of the central metabolic pathways. ''Metab Eng.'' '''5:''' 133-149 |
# Yoshida K, Yamaguchi M, Morinaga T, Kinehara M, Ikeuchi M, Ashida H, Fujita Y. (2008) myo-Inositol catabolism in Bacillus subtilis. ''J Biol Chem. '' ''' Apr 18;283(16):''' 10415-24. [http://www.ncbi.nlm.nih.gov/sites/entrez/18310071 PubMed] | # Yoshida K, Yamaguchi M, Morinaga T, Kinehara M, Ikeuchi M, Ashida H, Fujita Y. (2008) myo-Inositol catabolism in Bacillus subtilis. ''J Biol Chem. '' ''' Apr 18;283(16):''' 10415-24. [http://www.ncbi.nlm.nih.gov/sites/entrez/18310071 PubMed] | ||
Revision as of 23:24, 2 April 2009
- Description: formation of 5-deoxy-D-glucuronic acid (3rd reaction)
| Gene name | iolD |
| Synonyms | yxdD |
| Essential | no |
| Product | formation of 5-deoxy-D-glucuronic acid (3rd reaction) |
| Function | myo-inositol catabolism |
| MW, pI | 63 kDa, 5.006 |
| Gene length, protein length | 1740 bp, 580 aa |
| Immediate neighbours | iolE, iolC |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 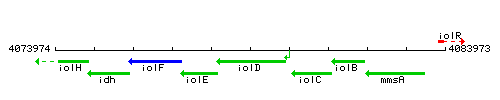
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor: SigA
- Regulation: repressed by glucose (CcpA)
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149
- Yoshida K, Yamaguchi M, Morinaga T, Kinehara M, Ikeuchi M, Ashida H, Fujita Y. (2008) myo-Inositol catabolism in Bacillus subtilis. J Biol Chem. Apr 18;283(16): 10415-24. PubMed