Difference between revisions of "AraE"
| Line 93: | Line 93: | ||
* '''Sigma factor:''' | * '''Sigma factor:''' | ||
| − | * '''Regulation:''' induced by arabinose [http://www.ncbi.nlm.nih.gov/sites/entrez/9620981 PubMed] | + | * '''Regulation:''' expression activated by glucose [http://www.ncbi.nlm.nih.gov/pubmed/12850135 PubMed], induced by arabinose [http://www.ncbi.nlm.nih.gov/sites/entrez/9620981 PubMed] |
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| Line 119: | Line 119: | ||
=References= | =References= | ||
| + | # Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in ''Bacillus subtilis'': regulation of the central metabolic pathways. ''Metab Eng.'' '''5:''' 133-149 [http://www.ncbi.nlm.nih.gov/pubmed/12850135 PubMed] | ||
# Krispin, O., and R. Allmansberger. (1998) The ''Bacillus subtilis'' AraE protein displays a broad substrate specificity for several different sugars. ''J. Bacteriol.'' '''180:''' 3250-3252. [http://www.ncbi.nlm.nih.gov/sites/entrez/9620981 PubMed] | # Krispin, O., and R. Allmansberger. (1998) The ''Bacillus subtilis'' AraE protein displays a broad substrate specificity for several different sugars. ''J. Bacteriol.'' '''180:''' 3250-3252. [http://www.ncbi.nlm.nih.gov/sites/entrez/9620981 PubMed] | ||
# Sá-Nogueira I, Ramos SS. (1997) Cloning, functional analysis, and transcriptional regulation of the Bacillus subtilis araE gene involved in L-arabinose utilization. ''J Bacteriol. '' '''Dec;179(24):''' 7705-11. [http://www.ncbi.nlm.nih.gov/sites/entrez/9401028 PubMed] | # Sá-Nogueira I, Ramos SS. (1997) Cloning, functional analysis, and transcriptional regulation of the Bacillus subtilis araE gene involved in L-arabinose utilization. ''J Bacteriol. '' '''Dec;179(24):''' 7705-11. [http://www.ncbi.nlm.nih.gov/sites/entrez/9401028 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 14:47, 2 April 2009
- Description: L-arabinose permease
| Gene name | araE |
| Synonyms | yvbR |
| Essential | no |
| Product | L-arabinose permease |
| Function | uptake of arabinose, galactose and xylose |
| MW, pI | 50 kDa, 9.259 |
| Gene length, protein length | 1392 bp, 464 aa |
| Immediate neighbours | cggR, araR |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 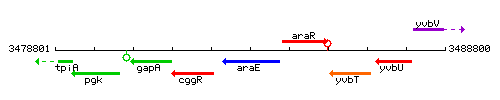
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transports arabinose, and also xylose and galactose PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149 PubMed
- Krispin, O., and R. Allmansberger. (1998) The Bacillus subtilis AraE protein displays a broad substrate specificity for several different sugars. J. Bacteriol. 180: 3250-3252. PubMed
- Sá-Nogueira I, Ramos SS. (1997) Cloning, functional analysis, and transcriptional regulation of the Bacillus subtilis araE gene involved in L-arabinose utilization. J Bacteriol. Dec;179(24): 7705-11. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed