Difference between revisions of "GudB"
| Line 20: | Line 20: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ypdA]]'', ''[[ypbH]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ypdA]]'', ''[[ypbH]]'' | ||
|- | |- | ||
| − | |style="background:#FAF8CC;" align="center"|'''[http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB14212]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
| − | |||
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:gudB_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:gudB_context.gif]] | ||
Revision as of 23:35, 16 April 2009
- Description: trigger enzyme: glutamate dehydrogenase (cryptic in 168 and derivatives)
| Gene name | gudB |
| Synonyms | ypcA |
| Essential | no |
| Product | glutamate dehydrogenase |
| Function | glutamate utilization, control of GltC activity |
| MW, pI | 47 kDa, 5.582 |
| Gene length, protein length | 1278 bp, 426 aa |
| Immediate neighbours | ypdA, ypbH |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 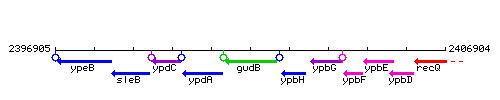
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
The gene is cryptic. If gudB is activated (gudB1 mutation), the bacteria are able to utilize glutamate as the only carbon source. PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s): RocG
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: gudB PubMed
- Regulation: constitutively expressed
- Regulatory mechanism:
- Additional information: GudB is subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant: GP691 (cat), available in Stülke lab
- Expression vector:
- GFP fusion:
- two-hybrid system:
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Belitsky BR, Sonenshein AL (1998) Role and regulation of Bacillus subtilis glutamate dehydrogenase genes. J Bacteriol 180:6298-6305 PubMed
- Commichau, F. M., Wacker, I., Schleider, J., Blencke, H.-M., Reif, I., Tripal, P., and Stülke, J. (2007) Characterization of Bacillus subtilis mutants with carbon source-independent glutamate biosynthesis. J Mol Microbiol Biotechnol 12: 106-113. PubMed
- Commichau, F. M., Gunka, K., Landmann, J. J. & Stülke, J. (2008) Glutamate metabolism in Bacillus subtilis: Gene expression and enzyme activities evolved to avoid futile cycles and to allow rapid responses to perturbations in the system. J. Bacteriol. 190: 3557-3564. PubMed
- Gerth et al. (2008) Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis. J Bacteriol 190:321-331 PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed