Difference between revisions of "YhaM"
| Line 26: | Line 26: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yhaM_1069042_1069986_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:yhaM_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yhaM_1069042_1069986_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:yhaM_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU09930]] |
|- | |- | ||
|} | |} | ||
Revision as of 12:53, 16 May 2013
- Description: RNase, 3'-> 5' exoribonuclease
| Gene name | yhaM |
| Synonyms | |
| Essential | no |
| Product | RNase YhaM |
| Function | RNA degradation |
| Gene expression levels in SubtiExpress: yhaM | |
| MW, pI | 35 kDa, 5.851 |
| Gene length, protein length | 942 bp, 314 aa |
| Immediate neighbours | sbcE, yhaL |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 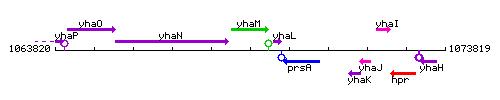
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed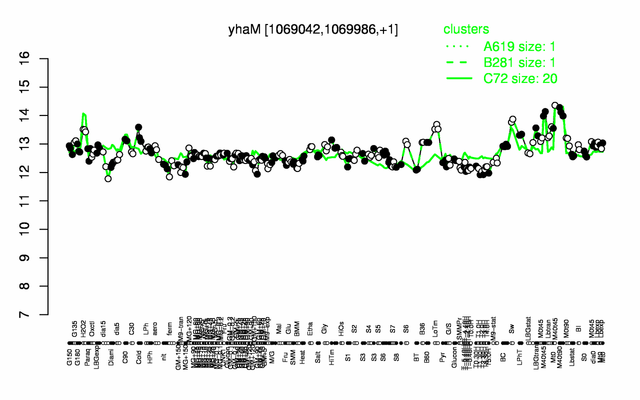
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU09930
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 3'-5' exoribonuclease
- Protein family: OB DNA-binding domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (Homogeneous) PubMed
Database entries
- Structure:
- UniProt: O07521
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Sigma factor:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
David Bechhofer, Mount Sinai School, New York, USA Homepage
Your additional remarks
References
Yulia Redko, Ciarán Condon
Maturation of 23S rRNA in Bacillus subtilis in the absence of Mini-III.
J Bacteriol: 2010, 192(1);356-9
[PubMed:19880604]
[WorldCat.org]
[DOI]
(I p)
Rita A Eckart, Sabine Brantl, Andreas Licht
Search for additional targets of the transcriptional regulator CcpN from Bacillus subtilis.
FEMS Microbiol Lett: 2009, 299(2);223-31
[PubMed:19732150]
[WorldCat.org]
[DOI]
(I p)
Ming Fang, Wencke-Maria Zeisberg, Ciaran Condon, Vasily Ogryzko, Antoine Danchin, Undine Mechold
Degradation of nanoRNA is performed by multiple redundant RNases in Bacillus subtilis.
Nucleic Acids Res: 2009, 37(15);5114-25
[PubMed:19553197]
[WorldCat.org]
[DOI]
(I p)
Ulf Gerth, Holger Kock, Ilja Kusters, Stephan Michalik, Robert L Switzer, Michael Hecker
Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis.
J Bacteriol: 2008, 190(1);321-31
[PubMed:17981983]
[WorldCat.org]
[DOI]
(I p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Nora Au, Elke Kuester-Schoeck, Veena Mandava, Laura E Bothwell, Susan P Canny, Karen Chachu, Sierra A Colavito, Shakierah N Fuller, Eli S Groban, Laura A Hensley, Theresa C O'Brien, Amish Shah, Jessica T Tierney, Louise L Tomm, Thomas M O'Gara, Alexi I Goranov, Alan D Grossman, Charles M Lovett
Genetic composition of the Bacillus subtilis SOS system.
J Bacteriol: 2005, 187(22);7655-66
[PubMed:16267290]
[WorldCat.org]
[DOI]
(P p)
Irina A Oussenko, Teppei Abe, Hiromi Ujiie, Akira Muto, David H Bechhofer
Participation of 3'-to-5' exoribonucleases in the turnover of Bacillus subtilis mRNA.
J Bacteriol: 2005, 187(8);2758-67
[PubMed:15805522]
[WorldCat.org]
[DOI]
(P p)
Irina A Oussenko, Roberto Sanchez, David H Bechhofer
Bacillus subtilis YhaM, a member of a new family of 3'-to-5' exonucleases in gram-positive bacteria.
J Bacteriol: 2002, 184(22);6250-9
[PubMed:12399495]
[WorldCat.org]
[DOI]
(P p)