Difference between revisions of "RnjA"
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || RNA processing | |style="background:#ABCDEF;" align="center"|'''Function''' || RNA processing | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU14530 rnjA] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/RnjA RNase J1] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/RnjA RNase J1] | ||
| Line 26: | Line 26: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[adeC]]'', ''[[ykzG]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[adeC]]'', ''[[ykzG]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU14530 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU14530 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU14530 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ykqC_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ykqC_context.gif]] | ||
Revision as of 12:43, 13 May 2013
- Description: RNase J1
| Gene name | rnjA |
| Synonyms | ykqC |
| Essential | yes PubMed |
| Product | RNase J1 |
| Function | RNA processing |
| Gene expression levels in SubtiExpress: rnjA | |
| Interactions involving this protein in SubtInteract: RNase J1 | |
| Metabolic function and regulation of this protein in SubtiPathways: Ile, Leu, Val, Coenzyme A | |
| MW, pI | 61 kDa, 5.902 |
| Gene length, protein length | 1665 bp, 555 aa |
| Immediate neighbours | adeC, ykzG |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 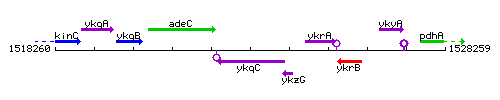
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed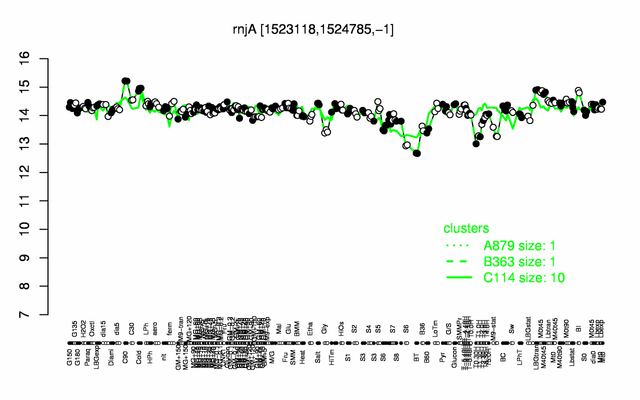
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU14530
Phenotypes of a mutant
- essential PubMed
- a study from the lab of Ciaran Condon reports that rnjA is non-essential and that the mutant is strongly impaired in sporulation, genetic competence and many other phenotypes PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: endonuclease and 5'-3' exonuclease
- Protein family: RNase J subfamily (according to Swiss-Prot)
- Paralogous protein(s): RnjB
RNAs affected by rnjA
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: Q45493
- KEGG entry: [2]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- required for thrS RNA processing, involved in maturation of the 5’-end of the16S rRNA
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant: GP41 (rnjA under control of p(xyl)), available in Stülke lab; SSB342 (rnjA under pspac), cat, available in Harald Putzer lab
- Expression vector:
- for chromosomal expression of RNase J1-Strep (spc): GP1034, available in Jörg Stülke's lab
- for chromosomal expression of RNase J1-Strep (cat): GP1042, available in Jörg Stülke's lab
- lacZ fusion: pGP418 (in pAC7), available in Jörg Stülke's lab
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- FLAG-tag construct:
- GP1020 (spc, based on pGP1331), available in Jörg Stülke's lab
- GP1075 (aphA3), available in Jörg Stülke's lab
- Antibody:
Labs working on this gene/protein
Harald Putzer, IBPC Paris, France Homepage
David Bechhofer, Mount Sinai School, New York, USA Homepage
Ciaran Condon, IBPC, Paris, France Homepage
Your additional remarks
References
Reviews
Zbigniew Dominski, Agamemnon J Carpousis, Béatrice Clouet-d'Orval
Emergence of the β-CASP ribonucleases: highly conserved and ubiquitous metallo-enzymes involved in messenger RNA maturation and degradation.
Biochim Biophys Acta: 2013, 1829(6-7);532-51
[PubMed:23403287]
[WorldCat.org]
[DOI]
(P p)
Martin Lehnik-Habrink, Richard J Lewis, Ulrike Mäder, Jörg Stülke
RNA degradation in Bacillus subtilis: an interplay of essential endo- and exoribonucleases.
Mol Microbiol: 2012, 84(6);1005-17
[PubMed:22568516]
[WorldCat.org]
[DOI]
(I p)
David H Bechhofer
Bacillus subtilis mRNA decay: new parts in the toolkit.
Wiley Interdiscip Rev RNA: 2011, 2(3);387-94
[PubMed:21957024]
[WorldCat.org]
[DOI]
(I p)
Jamie Richards, Joel G Belasco
Ribonuclease J: how to lead a double life.
Structure: 2011, 19(9);1201-3
[PubMed:21893280]
[WorldCat.org]
[DOI]
(I p)
Ciarán Condon, David H Bechhofer
Regulated RNA stability in the Gram positives.
Curr Opin Microbiol: 2011, 14(2);148-54
[PubMed:21334965]
[WorldCat.org]
[DOI]
(I p)
Ciarán Condon
What is the role of RNase J in mRNA turnover?
RNA Biol: 2010, 7(3);316-21
[PubMed:20458164]
[WorldCat.org]
[DOI]
(I p)
Original publications