Difference between revisions of "MaeA"
(→References) |
|||
| Line 141: | Line 141: | ||
=References= | =References= | ||
| + | <pubmed> 23136871</pubmed> | ||
<big>''Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J'' </big> | <big>''Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J'' </big> | ||
<big>'''RNA processing in ''Bacillus subtilis'': identification of targets of the essential RNase Y.''' </big> | <big>'''RNA processing in ''Bacillus subtilis'': identification of targets of the essential RNase Y.''' </big> | ||
Revision as of 11:05, 10 November 2012
- Description: malic enzyme
| Gene name | maeA |
| Synonyms | ywkA |
| Essential | no |
| Product | NAD-dependent malate dehydrogenase |
| Function | malate utilization |
| Gene expression levels in SubtiExpress: maeA | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 63 kDa, 4.883 |
| Gene length, protein length | 1746 bp, 582 aa |
| Immediate neighbours | ywkB, tdk |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 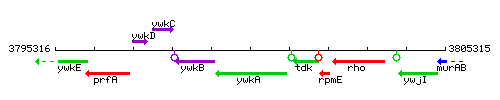
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed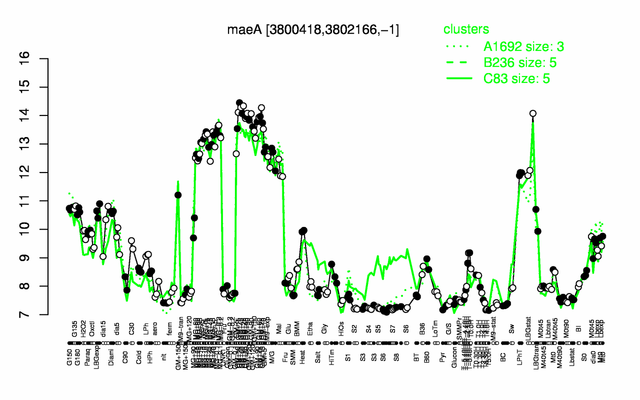
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU37050
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (S)-malate + NAD+ = pyruvate + CO2 + NADH (according to Swiss-Prot) malate --> pyruvate
- Protein family: malic enzymes family (according to Swiss-Prot)
- Paralogous protein(s): MalS
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P45868
- KEGG entry: [3]
- E.C. number: 1.1.1.38
Additional information
Expression and regulation
Biological materials
- Mutant: GP1141 (aphA3) available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Your additional remarks
References
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Guillaume Lerondel, Thierry Doan, Nicola Zamboni, Uwe Sauer, Stéphane Aymerich
YtsJ has the major physiological role of the four paralogous malic enzyme isoforms in Bacillus subtilis.
J Bacteriol: 2006, 188(13);4727-36
[PubMed:16788182]
[WorldCat.org]
[DOI]
(P p)
Thierry Doan, Pascale Servant, Shigeo Tojo, Hirotake Yamaguchi, Guillaume Lerondel, Ken-Ichi Yoshida, Yasutaro Fujita, Stéphane Aymerich
The Bacillus subtilis ywkA gene encodes a malic enzyme and its transcription is activated by the YufL/YufM two-component system in response to malate.
Microbiology (Reading): 2003, 149(Pt 9);2331-2343
[PubMed:12949160]
[WorldCat.org]
[DOI]
(P p)