Difference between revisions of "ClpP"
(→Targets of ClpC-ClpP-dependent protein degradation) |
(→Targets of ClpC-ClpP-dependent protein degradation) |
||
| Line 83: | Line 83: | ||
=== Targets of [[ClpC]]-[[ClpP]]-dependent protein degradation === | === Targets of [[ClpC]]-[[ClpP]]-dependent protein degradation === | ||
| − | * | + | * [[DegU]]-P {{PubMed|20070525}}, [[MurAA]] {{PubMed|14763982}}, [[ComK]] {{PubMed|9890793}}, [[CtsR]] {{PubMed|17380125}}, [[Mdh]], [[MgsR]] {{PubMed|22812682}} |
| + | |||
| + | === Targets of [[ClpX]]-[[ClpP]]-dependent protein degradation === | ||
| + | * [[MgsR]] {{PubMed|22812682}} | ||
=== Extended information on the protein === | === Extended information on the protein === | ||
Revision as of 16:43, 23 July 2012
- Description: ATP-dependent Clp protease proteolytic subunit (class III heat-shock protein)
| Gene name | clpP |
| Synonyms | yvdN |
| Essential | no |
| Product | ATP-dependent Clp protease proteolytic subunit |
| Function | protein degradation |
| Interactions involving this protein in SubtInteract: ClpP | |
| Metabolic function and regulation of this protein in SubtiPathways: Phosphorelay, Stress | |
| MW, pI | 21 kDa, 5.008 |
| Gene length, protein length | 591 bp, 197 aa |
| Immediate neighbours | trnQ-Arg, pgcM |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 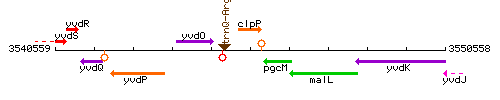
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed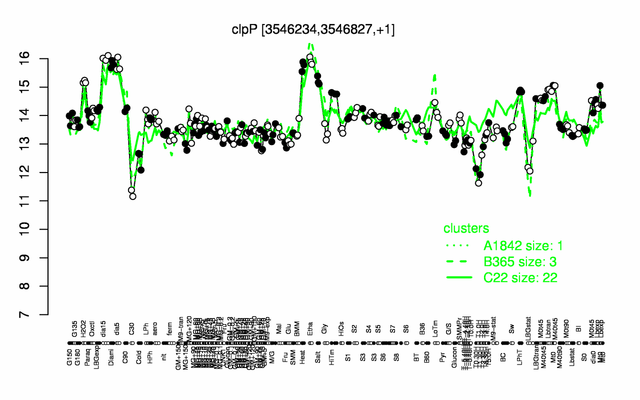
| |
Contents
Categories containing this gene/protein
proteolysis, general stress proteins (controlled by SigB), heat shock proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU34540
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Hydrolysis of proteins to small peptides in the presence of ATP and magnesium (according to Swiss-Prot) endopeptidase/proteolysis
- Protein family: peptidase S14 family (according to Swiss-Prot) ClpP (IPR001907) InterPro, (PF00574) PFAM
- Paralogous protein(s):
Targets of ClpC-ClpP-dependent protein degradation
Targets of ClpX-ClpP-dependent protein degradation
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-13 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- UniProt: P80244
- KEGG entry: [3]
- E.C. number: 3.4.21.92
Additional information
Expression and regulation
- Operon: clpP PubMed
- Additional information:
Biological materials
- Mutant:
- clpP::spec and clpP::cat, available in the Leendert Hamoen lab
- GP551 (spc), available in the Stülke lab
- Expression vector:
- lacZ fusion:
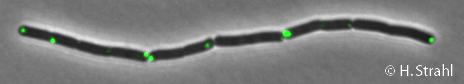
- GFP fusion: C-terminal GFP fusions (both single copy and as 2th copy in amyE locus, also as CFP and YFP variants) available in the Leendert Hamoen lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Newcastle University, UK homepage
Your additional remarks
References
Reviews
Additional reviews: PubMed
Original Publications
Additional publications: PubMed