Difference between revisions of "ComEB"
(→Expression and regulation) |
|||
| Line 90: | Line 90: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[comEA]]-[[comEB]]-[[comEC]]'' | + | * '''Operon:''' ''[[comEA]]-[[comEB]]-[[comEC]]'' {{PubMed|7968523,19028902}} |
| − | * '''[[Sigma factor]]:''' | + | * '''[[Sigma factor]]:''' [[SigA]] {{PubMed|7968523}} |
* '''Regulation:''' | * '''Regulation:''' | ||
| − | * '''Regulatory mechanism:''' post-translational control by [[ComN]] | + | * '''Regulatory mechanism:''' |
| + | ** post-translational control by [[ComN]] {{PubMed|19028902}} | ||
| + | ** ''[[yutB]]'' is required for transcription {{PubMed|19028902}} | ||
| + | ** [[ComK]]: transcription activation {{PubMed|8196543}} | ||
* '''Additional information:''' | * '''Additional information:''' | ||
Revision as of 18:49, 22 November 2009
- Description: late competence protein required for DNA binding and uptake
| Gene name | comEB |
| Synonyms | comD |
| Essential | no |
| Product | late competence protein |
| Function | genetic compentence |
| Metabolic function and regulation of this protein in SubtiPathways: Nucleotides (regulation) | |
| MW, pI | 20 kDa, 5.368 |
| Gene length, protein length | 567 bp, 189 aa |
| Immediate neighbours | comEC, comEA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 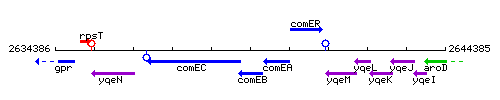
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU25580
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: cytidine and deoxycytidylate deaminase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P32393
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References