Difference between revisions of "SwrAA/1"
| Line 92: | Line 92: | ||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
| − | ** [[SwrA]]-[[DegU]] (N-terminal domain) {{PubMed|22496484}} | + | ** [[SwrA]]-[[DegU]]-P (N-terminal domain) {{PubMed|24386445,22496484}} |
* '''[[Localization]]:''' cytoplasm (according to Swiss-Prot) | * '''[[Localization]]:''' cytoplasm (according to Swiss-Prot) | ||
| Line 144: | Line 144: | ||
==Reviews== | ==Reviews== | ||
<pubmed>20735481 22092493 </pubmed> | <pubmed>20735481 22092493 </pubmed> | ||
| − | |||
==Original publications== | ==Original publications== | ||
| − | <pubmed>16091050 22773650 16357223,19389763,16030230,18567663,15066026 22496484 19389763 19749039 21278284 21602220,22329926,23190039</pubmed> | + | <pubmed>16091050 22773650 16357223,19389763,16030230,18567663,15066026 22496484 19389763 19749039 21278284 21602220,22329926,23190039 24386445 </pubmed> |
[[Category:Pseudogenes]] | [[Category:Pseudogenes]] | ||
Revision as of 14:46, 29 December 2014
- Description: control of DegU activity, enhances sigD transcription, controls the number of flagellar basal bodies, inactive pseudogene in strain 168
| Gene name | swrAA/1 |
| Synonyms | yvzD, swrAA, ifm |
| Essential | no |
| Product | swarming motility protein |
| Function | control of DegU activity |
| Gene expression levels in SubtiExpress: swrAA/1 | |
| Interactions involving this protein in SubtInteract: SwrA | |
| MW, pI | - , - |
| Gene length, protein length | 336 bp, - |
| Immediate neighbours | minJ, swrAA/2 |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 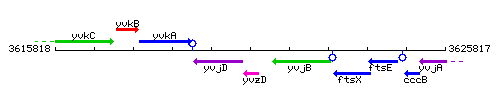
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed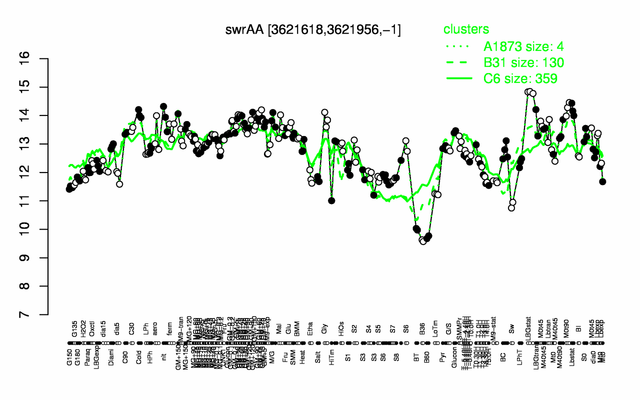
| |
Contents
Categories containing this gene/protein
biofilm formation, motility and chemotaxis, transcription factors and their control, pseudogenes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU35230
Phenotypes of a mutant
Database entries
- BsubCyc: BSU35230
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
- This protein is functional in undomesticated strains of B.subtilis but not in laboratory strains, such as 168, because of a frameshift mutation. Therefore laboratory strains of B.subtilis are unable to swarm.
- The frameshift in strain 168 is caused by a single insertion of an adenine in the codon for Tyr-12 which leads to the premature truncation of the protein in residue 13. In addition, the C-terminal section of swrAA was predicted to be an ORF (yvzD) by the genome project.
- Correction of sfp, epsC, swrAA, and degQ as well as introduction of rapP from a plasmid present in NCIB3610 results in biofilm formation in B. subtilis 168 PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU35230
- Structure:
- Swiss prot entry: O32266
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Serena Mordini, Cecilia Osera, Simone Marini, Francesco Scavone, Riccardo Bellazzi, Alessandro Galizzi, Cinzia Calvio
The role of SwrA, DegU and P(D3) in fla/che expression in B. subtilis.
PLoS One: 2013, 8(12);e85065
[PubMed:24386445]
[WorldCat.org]
[DOI]
(I e)
Sarah B Guttenplan, Sidney Shaw, Daniel B Kearns
The cell biology of peritrichous flagella in Bacillus subtilis.
Mol Microbiol: 2013, 87(1);211-29
[PubMed:23190039]
[WorldCat.org]
[DOI]
(I p)
Emilia Ghelardi, Sara Salvetti, Mara Ceragioli, Sokhna A Gueye, Francesco Celandroni, Sonia Senesi
Contribution of surfactin and SwrA to flagellin expression, swimming, and surface motility in Bacillus subtilis.
Appl Environ Microbiol: 2012, 78(18);6540-4
[PubMed:22773650]
[WorldCat.org]
[DOI]
(I p)
Mitsuo Ogura, Kensuke Tsukahara
SwrA regulates assembly of Bacillus subtilis DegU via its interaction with N-terminal domain of DegU.
J Biochem: 2012, 151(6);643-55
[PubMed:22496484]
[WorldCat.org]
[DOI]
(I p)
Loralyn M Cozy, Andrew M Phillips, Rebecca A Calvo, Ashley R Bate, Yi-Huang Hsueh, Richard Bonneau, Patrick Eichenberger, Daniel B Kearns
SlrA/SinR/SlrR inhibits motility gene expression upstream of a hypersensitive and hysteretic switch at the level of σ(D) in Bacillus subtilis.
Mol Microbiol: 2012, 83(6);1210-28
[PubMed:22329926]
[WorldCat.org]
[DOI]
(I p)
Kassem Hamze, Sabine Autret, Krzysztof Hinc, Soumaya Laalami, Daria Julkowska, Romain Briandet, Margareth Renault, Cédric Absalon, I Barry Holland, Harald Putzer, Simone J Séror
Single-cell analysis in situ in a Bacillus subtilis swarming community identifies distinct spatially separated subpopulations differentially expressing hag (flagellin), including specialized swarmers.
Microbiology (Reading): 2011, 157(Pt 9);2456-2469
[PubMed:21602220]
[WorldCat.org]
[DOI]
(I p)
Anna L McLoon, Sarah B Guttenplan, Daniel B Kearns, Roberto Kolter, Richard Losick
Tracing the domestication of a biofilm-forming bacterium.
J Bacteriol: 2011, 193(8);2027-34
[PubMed:21278284]
[WorldCat.org]
[DOI]
(I p)
Joyce E Patrick, Daniel B Kearns
Laboratory strains of Bacillus subtilis do not exhibit swarming motility.
J Bacteriol: 2009, 191(22);7129-33
[PubMed:19749039]
[WorldCat.org]
[DOI]
(I p)
Cecilia Osera, Giuseppe Amati, Cinzia Calvio, Alessandro Galizzi
SwrAA activates poly-gamma-glutamate synthesis in addition to swarming in Bacillus subtilis.
Microbiology (Reading): 2009, 155(Pt 7);2282-2287
[PubMed:19389763]
[WorldCat.org]
[DOI]
(P p)
Cinzia Calvio, Cecilia Osera, Giuseppe Amati, Alessandro Galizzi
Autoregulation of swrAA and motility in Bacillus subtilis.
J Bacteriol: 2008, 190(16);5720-8
[PubMed:18567663]
[WorldCat.org]
[DOI]
(I p)
Daniel B Kearns, Richard Losick
Cell population heterogeneity during growth of Bacillus subtilis.
Genes Dev: 2005, 19(24);3083-94
[PubMed:16357223]
[WorldCat.org]
[DOI]
(P p)
Nicola R Stanley, Beth A Lazazzera
Defining the genetic differences between wild and domestic strains of Bacillus subtilis that affect poly-gamma-dl-glutamic acid production and biofilm formation.
Mol Microbiol: 2005, 57(4);1143-58
[PubMed:16091050]
[WorldCat.org]
[DOI]
(P p)
Cinzia Calvio, Francesco Celandroni, Emilia Ghelardi, Giuseppe Amati, Sara Salvetti, Fabrizio Ceciliani, Alessandro Galizzi, Sonia Senesi
Swarming differentiation and swimming motility in Bacillus subtilis are controlled by swrA, a newly identified dicistronic operon.
J Bacteriol: 2005, 187(15);5356-66
[PubMed:16030230]
[WorldCat.org]
[DOI]
(P p)
Daniel B Kearns, Frances Chu, Rivka Rudner, Richard Losick
Genes governing swarming in Bacillus subtilis and evidence for a phase variation mechanism controlling surface motility.
Mol Microbiol: 2004, 52(2);357-69
[PubMed:15066026]
[WorldCat.org]
[DOI]
(P p)