Difference between revisions of "DacF"
| Line 120: | Line 120: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 95 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 10:07, 17 April 2014
- Description: penicillin-binding protein I, D-alanyl-D-alanine carboxypeptidase
| Gene name | dacF |
| Synonyms | |
| Essential | no |
| Product | penicillin-binding protein I, D-alanyl-D-alanine carboxypeptidase |
| Function | carboxypeptidase |
| Gene expression levels in SubtiExpress: dacF | |
| Metabolic function and regulation of this protein in SubtiPathways: dacF | |
| MW, pI | 43 kDa, 9.912 |
| Gene length, protein length | 1167 bp, 389 aa |
| Immediate neighbours | spoIIAA, pupG |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 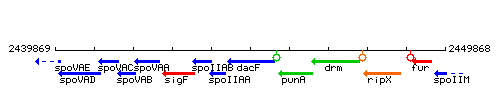
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed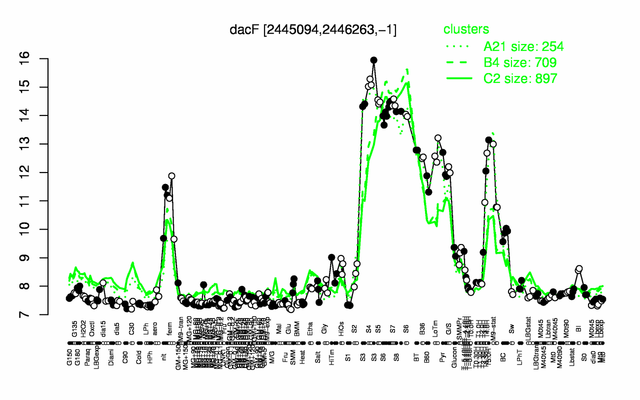
| |
Contents
Categories containing this gene/protein
cell wall synthesis, sporulation proteins
This gene is a member of the following regulons
SigF regulon, SigG regulon, AbrB regulon
The gene
Basic information
- Locus tag: BSU23480
Phenotypes of a mutant
Database entries
- BsubCyc: BSU23480
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: modifies degree of cross-linking of glycan strands in peptidoglycan
- Protein family: peptidase S11 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU23480
- Structure:
- UniProt: P38422
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
- number of protein molecules per cell (complex medium with amino acids, without glucose): 95 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Pengfei Zhang, Stacy Thomas, Yong-qing Li, Peter Setlow
Effects of cortex peptidoglycan structure and cortex hydrolysis on the kinetics of Ca(2+)-dipicolinic acid release during Bacillus subtilis spore germination.
J Bacteriol: 2012, 194(3);646-52
[PubMed:22123250]
[WorldCat.org]
[DOI]
(I p)
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Stephanie T Wang, Barbara Setlow, Erin M Conlon, Jessica L Lyon, Daisuke Imamura, Tsutomu Sato, Peter Setlow, Richard Losick, Patrick Eichenberger
The forespore line of gene expression in Bacillus subtilis.
J Mol Biol: 2006, 358(1);16-37
[PubMed:16497325]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
D L Popham, M E Gilmore, P Setlow
Roles of low-molecular-weight penicillin-binding proteins in Bacillus subtilis spore peptidoglycan synthesis and spore properties.
J Bacteriol: 1999, 181(1);126-32
[PubMed:9864321]
[WorldCat.org]
[DOI]
(P p)
Peter J Lewis, Jeffery Errington
Use of green fluorescent protein for detection of cell-specific gene expression and subcellular protein localization during sporulation in Bacillus subtilis.
Microbiology (Reading): 1996, 142 ( Pt 4);733-740
[PubMed:8936302]
[WorldCat.org]
[DOI]
(P p)