Difference between revisions of "PurL"
| Line 17: | Line 17: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=PurL PurL] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=PurL PurL] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/ | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=purL purL]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 80 kDa, 4.554 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 80 kDa, 4.554 | ||
Revision as of 10:17, 7 January 2014
- Description: phosphoribosylformylglycinamidine synthase
| Gene name | purL |
| Synonyms | |
| Essential | no |
| Product | phosphoribosylformylglycinamidine synthase |
| Function | purine biosynthesis |
| Gene expression levels in SubtiExpress: purL | |
| Interactions involving this protein in SubtInteract: PurL | |
| Metabolic function and regulation of this protein in SubtiPathways: purL | |
| MW, pI | 80 kDa, 4.554 |
| Gene length, protein length | 2226 bp, 742 aa |
| Immediate neighbours | purQ, purF |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 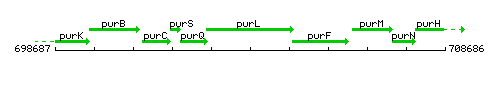
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed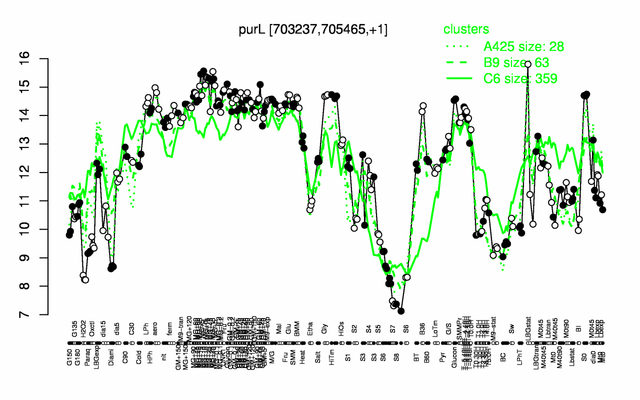
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of nucleotides, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU06480
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + N(2)-formyl-N(1)-(5-phospho-D-ribosyl)glycinamide + L-glutamine + H2O = ADP + phosphate + 2-(formamido)-N(1)-(5-phospho-D-ribosyl)acetamidine + L-glutamate (according to Swiss-Prot)
- Protein family: FGAMS family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot), membrane associated PubMed
Database entries
- UniProt: P12042
- KEGG entry: [3]
- E.C. number: 6.3.5.3
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Aaron A Hoskins, Ruchi Anand, Steven E Ealick, JoAnne Stubbe
The formylglycinamide ribonucleotide amidotransferase complex from Bacillus subtilis: metabolite-mediated complex formation.
Biochemistry: 2004, 43(32);10314-27
[PubMed:15301530]
[WorldCat.org]
[DOI]
(P p)
Lars Engholm Johansen, Per Nygaard, Catharina Lassen, Yvonne Agersø, Hans H Saxild
Definition of a second Bacillus subtilis pur regulon comprising the pur and xpt-pbuX operons plus pbuG, nupG (yxjA), and pbuE (ydhL).
J Bacteriol: 2003, 185(17);5200-9
[PubMed:12923093]
[WorldCat.org]
[DOI]
(P p)
M Weng, P L Nagy, H Zalkin
Identification of the Bacillus subtilis pur operon repressor.
Proc Natl Acad Sci U S A: 1995, 92(16);7455-9
[PubMed:7638212]
[WorldCat.org]
[DOI]
(P p)
D J Ebbole, H Zalkin
Cloning and characterization of a 12-gene cluster from Bacillus subtilis encoding nine enzymes for de novo purine nucleotide synthesis.
J Biol Chem: 1987, 262(17);8274-87
[PubMed:3036807]
[WorldCat.org]
(P p)