Difference between revisions of "Phy"
(→References) |
|||
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || untilization of inositol hexakisphosphate (phytate) | |style="background:#ABCDEF;" align="center"|'''Function''' || untilization of inositol hexakisphosphate (phytate) | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU19800 phy] |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 41 kDa, 4.936 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 41 kDa, 4.936 | ||
| Line 22: | Line 22: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[cgeB]]'', ''[[yodU]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[cgeB]]'', ''[[yodU]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU19800 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU19800 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU19800 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:phy_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:phy_context.gif]] | ||
Revision as of 13:02, 13 May 2013
- Description: phytase
| Gene name | phy |
| Synonyms | yodV, yzxA |
| Essential | no |
| Product | 3-phytase |
| Function | untilization of inositol hexakisphosphate (phytate) |
| Gene expression levels in SubtiExpress: phy | |
| MW, pI | 41 kDa, 4.936 |
| Gene length, protein length | 1146 bp, 382 aa |
| Immediate neighbours | cgeB, yodU |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 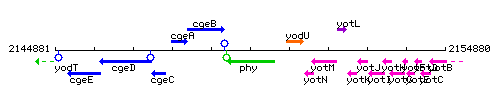
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed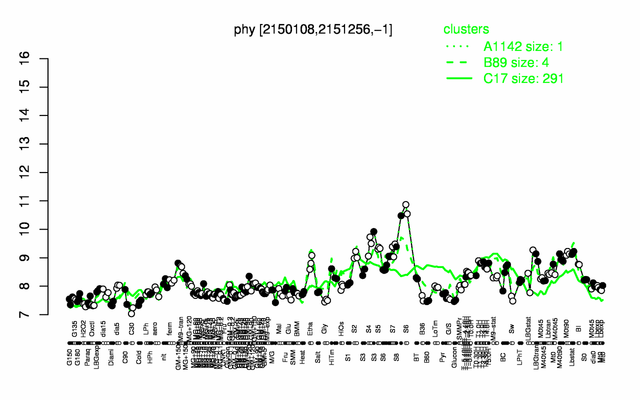
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU19800
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Myo-inositol hexakisphosphate + H2O = 1D-myo-inositol 1,2,4,5,6-pentakisphosphate + phosphate (according to Swiss-Prot)
- Protein family: msrA Met sulfoxide reductase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- extracellular (signal peptide) PubMed
Database entries
- Structure: 3AMR
- UniProt: P42094
- KEGG entry: [3]
- E.C. number: 3.1.3.8
Additional information
Expression and regulation
- Operon: phy PubMed
- Sigma factor: none PubMed
- Regulation:
- Additional information:
- the gene is not expressed in B. subtilis due to the absene of a functional promoter PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Martha Guerrero-Olazarán, Lilí Rodríguez-Blanco, J Gerardo Carreon-Treviño, Juan A Gallegos-López, José M Viader-Salvadó
Expression of a Bacillus phytase C gene in Pichia pastoris and properties of the recombinant enzyme.
Appl Environ Microbiol: 2010, 76(16);5601-8
[PubMed:20601512]
[WorldCat.org]
[DOI]
(I p)
Birgit Voigt, Haike Antelmann, Dirk Albrecht, Armin Ehrenreich, Karl-Heinz Maurer, Stefan Evers, Gerhard Gottschalk, Jan Maarten van Dijl, Thomas Schweder, Michael Hecker
Cell physiology and protein secretion of Bacillus licheniformis compared to Bacillus subtilis.
J Mol Microbiol Biotechnol: 2009, 16(1-2);53-68
[PubMed:18957862]
[WorldCat.org]
[DOI]
(I p)
Oliwia Makarewicz, Sarah Dubrac, Tarek Msadek, Rainer Borriss
Dual role of the PhoP approximately P response regulator: Bacillus amyloliquefaciens FZB45 phytase gene transcription is directed by positive and negative interactions with the phyC promoter.
J Bacteriol: 2006, 188(19);6953-65
[PubMed:16980498]
[WorldCat.org]
[DOI]
(P p)
N C Ha, B C Oh, S Shin, H J Kim, T K Oh, Y O Kim, K Y Choi, B H Oh
Crystal structures of a novel, thermostable phytase in partially and fully calcium-loaded states.
Nat Struct Biol: 2000, 7(2);147-53
[PubMed:10655618]
[WorldCat.org]
[DOI]
(P p)
S Roels, R Losick
Adjacent and divergently oriented operons under the control of the sporulation regulatory protein GerE in Bacillus subtilis.
J Bacteriol: 1995, 177(21);6263-75
[PubMed:7592393]
[WorldCat.org]
[DOI]
(P p)