Difference between revisions of "NtdA"
| Line 9: | Line 9: | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || pyridoxal phosphate-dependent 3-oxo-glucose-6-phosphate:glutamate aminotransferase | + | |style="background:#ABCDEF;" align="center"| '''Product''' || pyridoxal phosphate-dependent |
| + | 3-oxo-glucose-6-phosphate:glutamate aminotransferase | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || synthesis of the antibiotic kanosamine | |style="background:#ABCDEF;" align="center"|'''Function''' || synthesis of the antibiotic kanosamine | ||
Revision as of 13:39, 19 April 2013
- Description: pyridoxal phosphate-dependent 3-oxo-glucose-6-phosphate:glutamate aminotransferase
| Gene name | ntdA |
| Synonyms | yhjL |
| Essential | no |
| Product | pyridoxal phosphate-dependent
3-oxo-glucose-6-phosphate:glutamate aminotransferase |
| Function | synthesis of the antibiotic kanosamine |
| Gene expression levels in SubtiExpress: ntdA | |
| MW, pI | 49 kDa, 5.971 |
| Gene length, protein length | 1323 bp, 441 aa |
| Immediate neighbours | ntdB, ntdR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 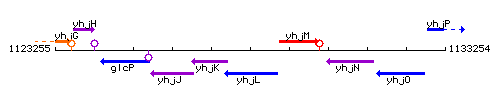
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed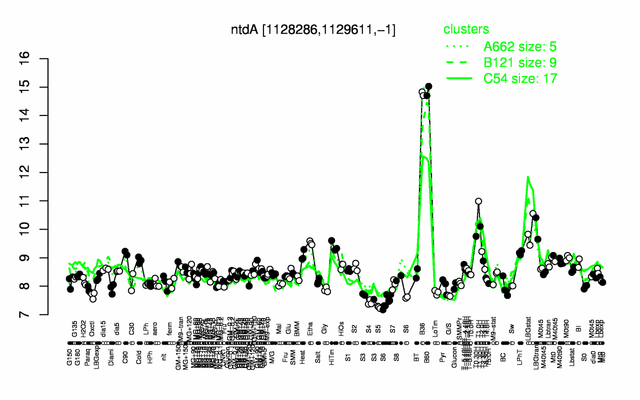
| |
Contents
Categories containing this gene/protein
miscellaneous metabolic pathways, biosynthesis of antibacterial compounds
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU10550
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- 3-oxo-d-glucose-6-phosphate + glutamate --> kanosamine-6-phosphate PubMed
- Protein family: degT/dnrJ/eryC1 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- pyridoxalphosphate PubMed
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O07566
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Natasha D Vetter, David M Langill, Shazia Anjum, Julie Boisvert-Martel, Rajendra C Jagdhane, Egiroh Omene, Hongyan Zheng, Karin E van Straaten, Isaac Asiamah, Ed S Krol, David A R Sanders, David R J Palmer
A previously unrecognized kanosamine biosynthesis pathway in Bacillus subtilis.
J Am Chem Soc: 2013, 135(16);5970-3
[PubMed:23586652]
[WorldCat.org]
[DOI]
(I p)
K E van Straaten, D M Langill, D R J Palmer, D A R Sanders
Purification, crystallization and preliminary X-ray analysis of NtdA, a putative pyridoxal phosphate-dependent aminotransferase from Bacillus subtilis.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2009, 65(Pt 4);426-9
[PubMed:19342798]
[WorldCat.org]
[DOI]
(I p)
Takashi Inaoka, Takenori Satomura, Yasutaro Fujita, Kozo Ochi
Novel gene regulation mediated by overproduction of secondary metabolite neotrehalosadiamine in Bacillus subtilis.
FEMS Microbiol Lett: 2009, 291(2);151-6
[PubMed:19087206]
[WorldCat.org]
[DOI]
(I p)
Takashi Inaoka, Kozo Ochi
Glucose uptake pathway-specific regulation of synthesis of neotrehalosadiamine, a novel autoinducer produced in Bacillus subtilis.
J Bacteriol: 2007, 189(1);65-75
[PubMed:17056753]
[WorldCat.org]
[DOI]
(P p)
Takashi Inaoka, Kosaku Takahashi, Hiroshi Yada, Mitsuru Yoshida, Kozo Ochi
RNA polymerase mutation activates the production of a dormant antibiotic 3,3'-neotrehalosadiamine via an autoinduction mechanism in Bacillus subtilis.
J Biol Chem: 2004, 279(5);3885-92
[PubMed:14612444]
[WorldCat.org]
[DOI]
(P p)