Difference between revisions of "CitB"
| Line 52: | Line 52: | ||
{{SubtiWiki regulon|[[FsrA regulon]]}} | {{SubtiWiki regulon|[[FsrA regulon]]}} | ||
| − | =The [[CitB regulon]]: ''[[feuA]]-[[feuB]]-[[feuC]]-[[ybbA]]''= | + | =The [[CitB regulon]]: ''[[feuA]]-[[feuB]]-[[feuC]]-[[ybbA]]'', ''[[citZ]]''= |
=The gene= | =The gene= | ||
| Line 61: | Line 61: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | glutamate auxotrophy and a defect in sporulation [http://www.ncbi.nlm.nih.gov/pubmed/9393699 PubMed] | + | * glutamate auxotrophy and a defect in sporulation [http://www.ncbi.nlm.nih.gov/pubmed/9393699 PubMed] |
=== Database entries === | === Database entries === | ||
| Line 77: | Line 77: | ||
* '''Catalyzed reaction/ biological activity:''' | * '''Catalyzed reaction/ biological activity:''' | ||
| − | ** Citrate = isocitrate | + | ** Citrate <=> isocitrate |
| − | ** Binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster | + | ** Binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster {{PubMed|23354745,10468622}} |
* '''Protein family:''' | * '''Protein family:''' | ||
| Line 175: | Line 175: | ||
==Original publications== | ==Original publications== | ||
'''Additional publications:''' {{PubMed|20817675}} | '''Additional publications:''' {{PubMed|20817675}} | ||
| − | <pubmed>18697947, 20097860,2118511,12591885,10656796,12591885,12850135 2413006 10656796 10468622 6143742 12591885 16395550 16923907 9642180 9393699 12591885 2105305,20933603 21099137 21446632 21821766 22389480 23139400</pubmed> | + | <pubmed>18697947, 20097860,2118511,12591885,10656796,12591885,12850135 2413006 10656796 10468622 6143742 12591885 16395550 16923907 9642180 9393699 12591885 2105305,20933603 21099137 21446632 21821766 22389480 23139400 23354745 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 09:48, 29 January 2013
- Description: trigger enzyme: aconitase and RNA binding protein
| Gene name | citB |
| Synonyms | |
| Essential | no |
| Product | trigger enzyme: aconitate hydratase (aconitase) |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: citB | |
| Interactions involving this protein in SubtInteract: CitB | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 99 kDa, 4.903 |
| Gene length, protein length | 2727 bp, 909 aa |
| Immediate neighbours | sspO, yneN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 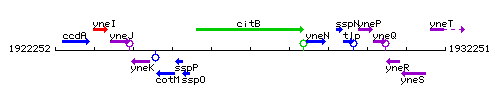
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed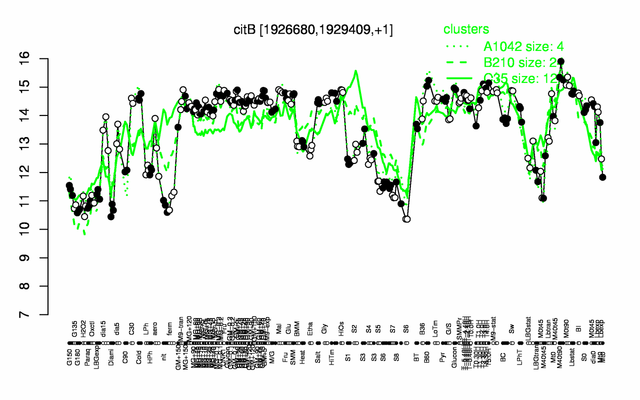
| |
Contents
Categories containing this gene/protein
carbon core metabolism, trigger enzyme, RNA binding regulators
This gene is a member of the following regulons
CcpA regulon, CcpC regulon, CodY regulon, FsrA regulon
The CitB regulon: feuA-feuB-feuC-ybbA, citZ
The gene
Basic information
- Locus tag: BSU18000
Phenotypes of a mutant
- glutamate auxotrophy and a defect in sporulation PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
- A mutation was found in this gene after evolution under relaxed selection for sporulation PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Citrate <=> isocitrate
- Binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): FeS cluster
- Effectors of protein activity:
Database entries
- Structure: 1L5J (E. coli)
- UniProt: P09339
- KEGG entry: [3]
- E.C. number: 4.2.1.3
Additional information
- B. subtilis aconitase is both an enzyme and an RNA binding protein (moonlighting protein) PubMed
- extensive information on the structure and enzymatic properties of CitB can be found at Proteopedia
Expression and regulation
- Regulation:
- repressed during growth in the presence of branched chain amino acids (CodY) PubMed
- repressed in the presence of glucose and glutamate (CcpC) PubMed
- expressed upon transition into the stationary phase (AbrB) PubMed, indirect negative regulation by AbrB PubMed
- repressed by glucose (3.7-fold) (CcpA) PubMed
- repression by glucose + arginine (CcpC) PubMed
- less expressed under conditions of extreme iron limitation (FsrA) PubMed
- part of the iron sparing response (FsrA) PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP683 (erm), available in Jörg Stülke's lab
- GP1441 (spc), available in Jörg Stülke's lab
- 1A999 ( citB::spec), PubMed, available at BGSC
- Expression vector:
- GP1439 (citB-Strep (spc)), purification from B. subtilis, for SPINE, available in Jörg Stülke's lab
- pGP1810 (for expression, purification in E. coli with N-terminal Strep-tag, in pGP172, available in Jörg Stülke's lab
- lacZ fusion:
- pGP700 (in pAC5), available in Jörg Stülke's lab
- GFP fusion: GP1434 (spc, based on pGP1870), available in Jörg Stülke's lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody: available in Linc Sonenshein's lab
- FLAG-tag construct:
- GP1144 (spc, based on pGP1331), available in Jörg Stülke's lab
- GP1145 (kan), available in Jörg Stülke's lab
Labs working on this gene/protein
- Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
- Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed