Difference between revisions of "PnpA"
(→Biological materials) |
(→Phenotypes of a mutant) |
||
| Line 45: | Line 45: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | * The ''pnpA'' mutant is cold sensitive and sensitive to tetracyclin, it shows multiseptate filamentous growth. | + | * The ''pnpA'' mutant is cold sensitive and sensitive to tetracyclin, it shows multiseptate filamentous growth. {{PubMed|8636041}} |
| − | * The mutant is deficient in genetic competence (no expression of the late competence genes) | + | * The mutant is deficient in genetic competence (no expression of the late competence genes) {{PubMed|8825779}} |
* The mutant overexpresses the ''trp'' and ''[[ycgM]]-[[ycgN]]-[[ycgO]]'' operons. | * The mutant overexpresses the ''trp'' and ''[[ycgM]]-[[ycgN]]-[[ycgO]]'' operons. | ||
Revision as of 12:37, 19 April 2011
- Description: polynucleotide phosphorylase, RNase
| Gene name | pnpA |
| Synonyms | comR |
| Essential | no |
| Product | polynucleotide phosphorylase (PNPase) (EC 2.7.7.8) |
| Function | necessary for competence development |
| MW, pI | 77 kDa, 4.89 |
| Gene length, protein length | 2115 bp, 705 aa |
| Immediate neighbours | rpsO, ylxY |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 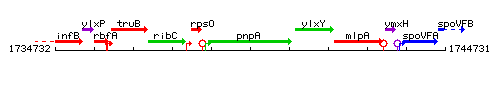
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU16690
Phenotypes of a mutant
- The pnpA mutant is cold sensitive and sensitive to tetracyclin, it shows multiseptate filamentous growth. PubMed
- The mutant is deficient in genetic competence (no expression of the late competence genes) PubMed
- The mutant overexpresses the trp and ycgM-ycgN-ycgO operons.
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 3'-5' exoribonuclease, RNase, PNPase degrades the trp mRNA from the RNA-TRAP complex
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions: part of the RNA degradosome
- Localization:
Database entries
- Structure: 3CDI (protein from E. coli), 3GCM (protein from E. coli, PNPase/RNase E micro-domain/RNA tetragonal crystal form )
- UniProt: P50849
- KEGG entry: [2]
- E.C. number:
Additional information
required for the expression of late competence genes comGA and comK, requirement bypassed by a mecA disruption; may be necessary for modification of the srfAA transcript (stabilization or translation activation)
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP584 (aphA3), available in Stülke lab
- Expression vector:
- for expression, purification in E. coli with N-terminal His-tag, in pWH844: pGP838, available in Stülke lab
- for expression/ purification from B. subtilis with N-terminal Strep-tag, for SPINE, in pGP380: pGP1342, available in Stülke lab
- for chromosomal expression of PnpA-Strep (cat): GP1002, available in Jörg Stülke's lab
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
David Bechhofer, Mount Sinai School, New York, USA Homepage
Your additional remarks
References
Reviews
Original publications