Difference between revisions of "LytB"
| Line 59: | Line 59: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU35630&redirect=T BSU35630] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/lytABC.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/lytABC.html] | ||
| Line 97: | Line 98: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU35630&redirect=T BSU35630] | ||
* '''Structure:''' | * '''Structure:''' | ||
Revision as of 14:57, 2 April 2014
- Description: modifier protein of major autolysin LytC
| Gene name | lytB |
| Synonyms | cwbA |
| Essential | no |
| Product | modifier protein of major autolysin LytC |
| Function | autolysis |
| Gene expression levels in SubtiExpress: lytB | |
| MW, pI | 76 kDa, 9.835 |
| Gene length, protein length | 2115 bp, 705 aa |
| Immediate neighbours | lytC, lytA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 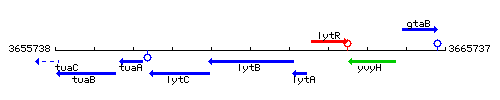
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed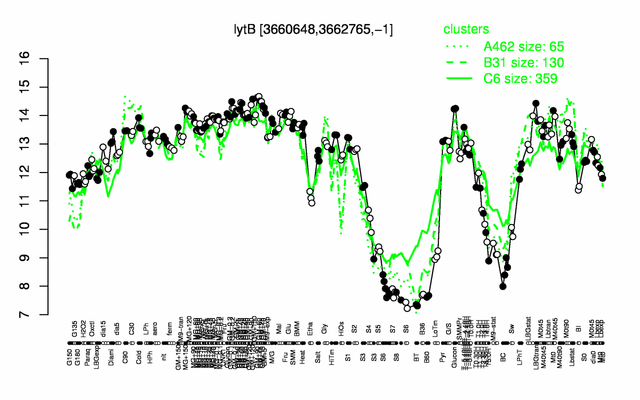
| |
Contents
Categories containing this gene/protein
cell wall degradation/ turnover, membrane proteins
This gene is a member of the following regulons
SigD regulon, SlrR regulon, YvrHb regulon
The gene
Basic information
- Locus tag: BSU35630
Phenotypes of a mutant
Database entries
- BsubCyc: BSU35630
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cell membrane (according to Swiss-Prot)
Database entries
- BsubCyc: BSU35630
- Structure:
- UniProt: Q02113
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Yunrong Chai, Thomas Norman, Roberto Kolter, Richard Losick
An epigenetic switch governing daughter cell separation in Bacillus subtilis.
Genes Dev: 2010, 24(8);754-65
[PubMed:20351052]
[WorldCat.org]
[DOI]
(I p)
Masakuni Serizawa, Keisuke Kodama, Hiroki Yamamoto, Kazuo Kobayashi, Naotake Ogasawara, Junichi Sekiguchi
Functional analysis of the YvrGHb two-component system of Bacillus subtilis: identification of the regulated genes by DNA microarray and northern blot analyses.
Biosci Biotechnol Biochem: 2005, 69(11);2155-69
[PubMed:16306698]
[WorldCat.org]
[DOI]
(P p)
P Margot, V Lazarevic, D Karamata
Effect of the SinR protein on the expression of the Bacillus subtilis 168 lytABC operon.
Microb Drug Resist: 1996, 2(1);119-21
[PubMed:9158733]
[WorldCat.org]
[DOI]
(P p)
V Lazarevic, P Margot, B Soldo, D Karamata
Sequencing and analysis of the Bacillus subtilis lytRABC divergon: a regulatory unit encompassing the structural genes of the N-acetylmuramoyl-L-alanine amidase and its modifier.
J Gen Microbiol: 1992, 138(9);1949-61
[PubMed:1357079]
[WorldCat.org]
[DOI]
(P p)
A Kuroda, J Sekiguchi
Characterization of the Bacillus subtilis CwbA protein which stimulates cell wall lytic amidases.
FEMS Microbiol Lett: 1992, 74(1);109-13
[PubMed:1355454]
[WorldCat.org]
[DOI]
(P p)