MinC
- Description: cell-division inhibitor (septum placement), destabilizes FtsZ-rings at new cell poles, part of the Min system (with DivIVA, MinD, MinJ), Noc and the Min system ensure the efficient utilization of the division site at midcell in by ensuring Z ring placement
| Gene name | minC |
| Synonyms | |
| Essential | no |
| Product | cell-division inhibitor |
| Function | septum placement |
| Gene expression levels in SubtiExpress: minC | |
| Interactions involving this protein in SubtInteract: MinC | |
| MW, pI | 24 kDa, 6.262 |
| Gene length, protein length | 678 bp, 226 aa |
| Immediate neighbours | minD, mreD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 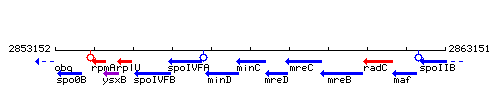
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed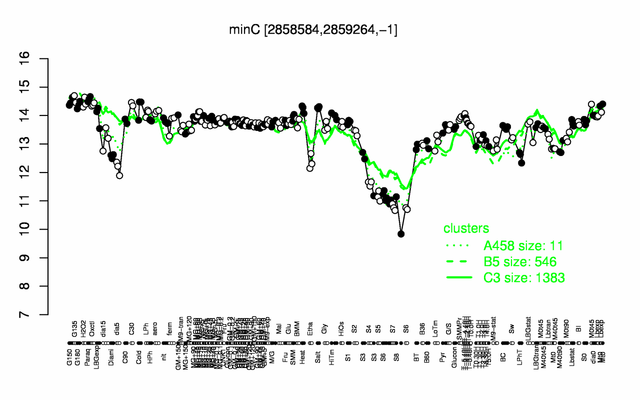
| |
Contents
Categories containing this gene/protein
cell division, cell envelope stress proteins (controlled by SigM, V, W, X, Y), membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28000
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28000
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: minC family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- BsubCyc: BSU28000
- Structure:
- UniProt: Q01463
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 95 PubMed
Biological materials
- Mutant: BSN375 (minC::aphA3) (available in Leendert Hamoen's, Sven Halbedel's and Jörg Stülke's labs)
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original Publications