YfhM
- Description: general stress protein, similar to epoxide hydrolase
| Gene name | yfhM |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | survival of ethanol stress |
| Gene expression levels in SubtiExpress: yfhM | |
| MW, pI | 32 kDa, 6.064 |
| Gene length, protein length | 858 bp, 286 aa |
| Immediate neighbours | yfhL, csbB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed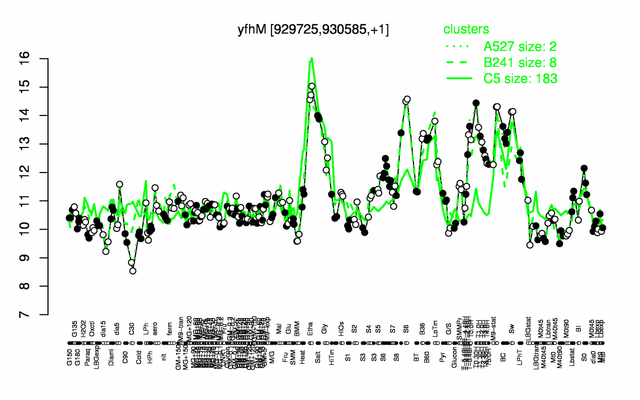
| |
Contents
Categories containing this gene/protein
general stress proteins (controlled by SigB), cell envelope stress proteins (controlled by SigM, V, W, X, Y), poorly characterized/ putative enzymes, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU08590
Phenotypes of a mutant
Database entries
- BsubCyc: BSU08590
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: AB hydrolase superfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU08590
- Structure:
- UniProt: O31581
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 194 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Dirk Höper, Uwe Völker, Michael Hecker
Comprehensive characterization of the contribution of individual SigB-dependent general stress genes to stress resistance of Bacillus subtilis.
J Bacteriol: 2005, 187(8);2810-26
[PubMed:15805528]
[WorldCat.org]
[DOI]
(P p)
H Antelmann, C Scharf, M Hecker
Phosphate starvation-inducible proteins of Bacillus subtilis: proteomics and transcriptional analysis.
J Bacteriol: 2000, 182(16);4478-90
[PubMed:10913081]
[WorldCat.org]
[DOI]
(P p)
X Huang, A Gaballa, M Cao, J D Helmann
Identification of target promoters for the Bacillus subtilis extracytoplasmic function sigma factor, sigma W.
Mol Microbiol: 1999, 31(1);361-71
[PubMed:9987136]
[WorldCat.org]
[DOI]
(P p)