LigD
- Description: DNA repair polymerase/ ligase in non-homologous end joining DNA repair
| Gene name | ligD |
| Synonyms | ykoU |
| Essential | no |
| Product | DNA repair polymerase/ ligase |
| Function | non-homologous end joining DNA repair, repair of gapped DNA substrates |
| Gene expression levels in SubtiExpress: ligD | |
| Interactions involving this protein in SubtInteract: LigD | |
| MW, pI | 70 kDa, 6.646 |
| Gene length, protein length | 1833 bp, 611 aa |
| Immediate neighbours | ykoT, ykoV |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 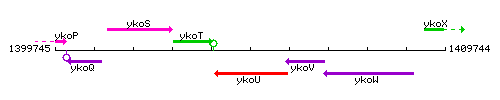
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed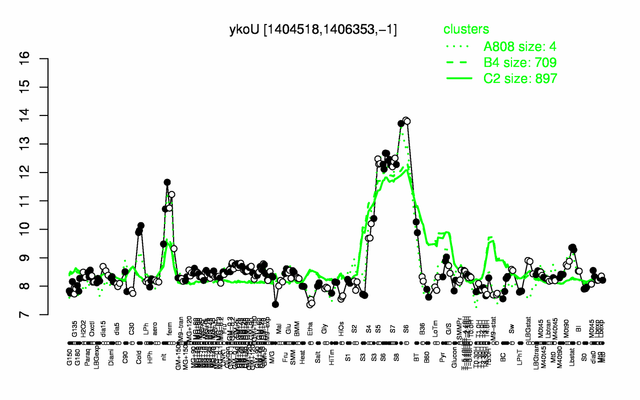
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13400
Phenotypes of a mutant
- sensitivity to ionizing radiation in the stationary phase PubMed
- sensitivity of spores to several DNA-damaging treatments known to cause double strand breaks, such as UV-ray, X-ray, ultrahigh vacuum and wet heat PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains: N-terminal DNA ligase catalytic domain (aa 1 - 331) linked to a C-terminal polymerase domain (aa 332 - 611) PubMed
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O34398
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- expressed during sporulation in the forespore (SigG, SpoVT) PubMed
- Additional information:
Biological materials
- Mutant:
- BP141 (ykoV-ligD::kan) available in Fabian Commichau's lab
- BP142 (ykoW-ykoV-ligD::kan) available in Fabian Commichau's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Justin S Lenhart, Jeremy W Schroeder, Brian W Walsh, Lyle A Simmons
DNA repair and genome maintenance in Bacillus subtilis.
Microbiol Mol Biol Rev: 2012, 76(3);530-64
[PubMed:22933559]
[WorldCat.org]
[DOI]
(I p)
Stewart Shuman, Michael S Glickman
Bacterial DNA repair by non-homologous end joining.
Nat Rev Microbiol: 2007, 5(11);852-61
[PubMed:17938628]
[WorldCat.org]
[DOI]
(I p)
Original publications