AroA
- Description: 3-deoxy-D-arabino-heptulosonate 7-phosphate synthase / chorismate mutase-isozyme 3
| Gene name | aroA |
| Synonyms | aroG |
| Essential | no |
| Product | 3-deoxy-D-arabino-heptulosonate 7-phosphate synthase /
chorismate mutase-isozyme 3 |
| Function | biosynthesis of aromatic amino acids |
| Metabolic function and regulation of this protein in SubtiPathways: Phe, Tyr, Trp | |
| MW, pI | 39 kDa, 5.341 |
| Gene length, protein length | 1074 bp, 358 aa |
| Immediate neighbours | ccpA, ytxJ |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 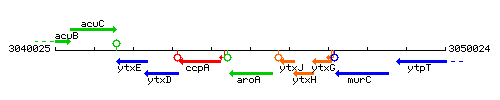
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed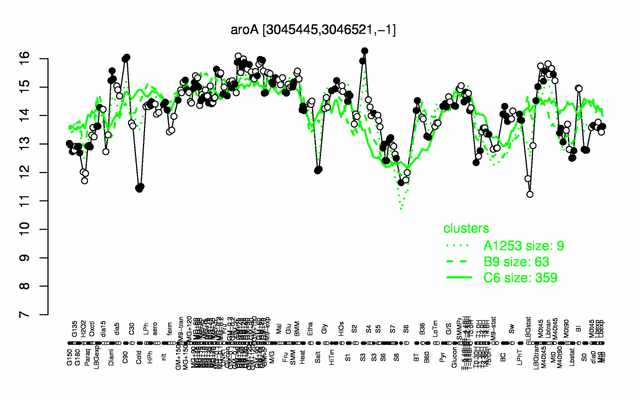
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29750
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Phosphoenolpyruvate + D-erythrose 4-phosphate + H2O = 3-deoxy-D-arabino-hept-2-ulosonate 7-phosphate + phosphate (according to Swiss-Prot)
- Protein family: class-I DAHP synthetase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity: subject to feedback inhibition PubMed
Database entries
- Structure: 1VR6 (from Thermotoga maritima, 51% identity, 68% similarity)
- UniProt: P39912
- KEGG entry: [3]
- E.C. number: 2.5.1.54 5 and 5.4.99.5
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information: subject to feedback inhibition PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Ana Gutiérrez-Preciado, Tina M Henkin, Frank J Grundy, Charles Yanofsky, Enrique Merino
Biochemical features and functional implications of the RNA-based T-box regulatory mechanism.
Microbiol Mol Biol Rev: 2009, 73(1);36-61
[PubMed:19258532]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Alia Lapidus, Nathalie Galleron, Alexei Sorokin, S Dusko Ehrlich
Sequencing and functional annotation of the Bacillus subtilis genes in the 200 kb rrnB-dnaB region.
Microbiology (Reading): 1997, 143 ( Pt 11);3431-3441
[PubMed:9387221]
[WorldCat.org]
[DOI]
(P p)