LysC
- Description: aspartokinase II (alpha and beta subunits)
| Gene name | lysC |
| Synonyms | ask, aecA |
| Essential | no |
| Product | aspartokinase II (alpha and beta subunits) |
| Function | biosynthesis of lysine |
| Metabolic function and regulation of this protein in SubtiPathways: Lys, Thr | |
| MW, pI | 43 kDa, 4.643 |
| Gene length, protein length | 1224 bp, 408 aa |
| Immediate neighbours | yslB, ask |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 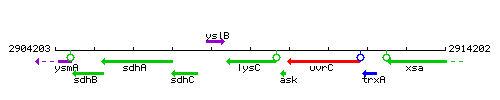
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed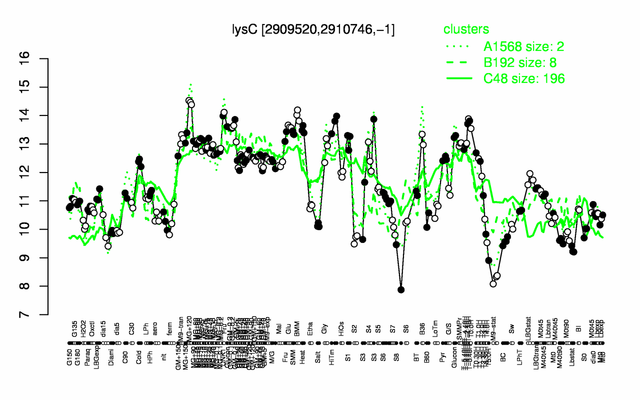
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28470
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + L-aspartate = ADP + 4-phospho-L-aspartate (according to Swiss-Prot)
- Protein family: aspartokinase family (according to Swiss-Prot)
- Paralogous protein(s): DapG
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2RE1 (from Neisseria meningitidis mc58, 40% identity, 58% similarity)
- UniProt: P08495
- KEGG entry: [3]
- E.C. number: 2.7.2.4
Additional information
Expression and regulation
- Operon: lysC PubMed
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed, also degraded upon ammonium or amino acid starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original Publications
The L-box riboswitch
Sharnise N Wilson-Mitchell, Frank J Grundy, Tina M Henkin
Analysis of lysine recognition and specificity of the Bacillus subtilis L box riboswitch.
Nucleic Acids Res: 2012, 40(12);5706-17
[PubMed:22416067]
[WorldCat.org]
[DOI]
(I p)
Simon Blouin, Raja Chinnappan, Daniel A Lafontaine
Folding of the lysine riboswitch: importance of peripheral elements for transcriptional regulation.
Nucleic Acids Res: 2011, 39(8);3373-87
[PubMed:21169337]
[WorldCat.org]
[DOI]
(I p)
Trang Thi Phuong Phan, Wolfgang Schumann
Transcriptional analysis of the lysine-responsive and riboswitch-regulated lysC gene of Bacillus subtilis.
Curr Microbiol: 2009, 59(4);463-8
[PubMed:19636616]
[WorldCat.org]
[DOI]
(I p)
Narasimhan Sudarsan, J Kenneth Wickiser, Shingo Nakamura, Margaret S Ebert, Ronald R Breaker
An mRNA structure in bacteria that controls gene expression by binding lysine.
Genes Dev: 2003, 17(21);2688-97
[PubMed:14597663]
[WorldCat.org]
[DOI]
(P p)
Frank J Grundy, Susan C Lehman, Tina M Henkin
The L box regulon: lysine sensing by leader RNAs of bacterial lysine biosynthesis genes.
Proc Natl Acad Sci U S A: 2003, 100(21);12057-62
[PubMed:14523230]
[WorldCat.org]
[DOI]
(P p)
Other original Publications