CcpA
- Description: Carbon catabolite control protein A, involved in glucose regulation of many genes; represses catabolic genes and activates genes involved in excretion of excess carbon
| Gene name | ccpA |
| Synonyms | graR, alsA, amyR |
| Essential | no |
| Product | transcriptional regulator (LacI family) |
| Function | mediates carbon catabolite repression (CCR) |
| Metabolic function and regulation of this protein in SubtiPathways: Nucleoside catabolism, Nucleotides (regulation), Ile, Leu, Val, His, Coenzyme A, Central C-metabolism | |
| MW, pI | 36,8 kDa, 5.06 |
| Gene length, protein length | 1002 bp, 334 amino acids |
| Immediate neighbours | motP, aroA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 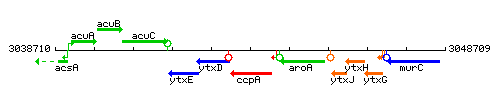
This image was kindly provided by SubtiList
| |
Contents
- 1 Categories containing this gene/protein
- 2 This gene is a member of the following regulons
- 3 The CcpA regulon
- 4 The gene
- 5 The protein
- 6 Expression and regulation
- 7 Biological materials
- 8 Labs working on this gene/protein
- 9 Your additional remarks
- 10 References
- 10.1 Reviews
- 10.2 General and physiological studies
- 10.3 Global analyses (proteome, transcriptome)
- 10.4 Repression of target genes by CcpA
- 10.5 Positive regulation of gene expression by CcpA
- 10.6 Control of CcpA activity
- 10.7 CcpA-DNA interaction
- 10.8 Functional analysis of CcpA
- 10.9 Structural analyses
Categories containing this gene/protein
This gene is a member of the following regulons
The CcpA regulon
The gene
Basic information
- Locus tag: BSU29740
Phenotypes of a mutant
Loss of carbon catabolite repression. Loss of PTS-dependent sugar transport due to excessive phosphorylation of HPr by HprK. The mutant is unable to grow on a minimal medium with glucose and ammonium as the only sources of carbon and nitrogen, respectively.
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transcriptional regulator of carbon catabolite repression (CCR)
- Protein family: LacI family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- HTH lacI-type Domain (1 – 58)
- DNA binding Domain (6 – 25)
- Modification:
- Effectors of protein activity:glucose-6-phosphate, fructose-1,6-bisphosphate Pubmed
- Localization:
Database entries
- Structure:
- 2JCG (Apoprotein from Bacillus megaterium)
- CcpA-Crh-DNA-complex NCBI
- complex with P-Ser-HPr and sulphate ions NCBI
- 3OQM (complex of B. subtilis CcpA with P-Ser-HPr and the ackA operator site)
- 3OQN (complex of B. subtilis CcpA with P-Ser-HPr and the gntR operator site)
- 3OQO (complex of B. subtilis CcpA with P-Ser-HPr and a optimal synthetic operator site)
- UniProt: P25144
- KEGG entry: [3]
Additional information
Expression and regulation
- Sigma factor:
- Regulation: constitutively expressed PubMed
- Additional information: there are about 3.000 molecules of CcpA per cell PubMed, this corresponds to a concentration of 3 myM (according to PubMed)
Biological materials
- Expression vector: pGP643 (N-terminal Strep-tag, purification from B. subtilis, for SPINE, in pGP380), available in Stülke lab
- lacZ fusion:
- GFP fusion:
Labs working on this gene/protein
- Wolfgang Hillen, Erlangen University, Germany Homepage
- Gerald Seidel, Erlangen University, Germany Homepage
- Richard Brennan, Houston, Texas, USA Homepage
- Milton H. Saier, University of California at San Diego, USA Homepage
- Yasutaro Fujita, University of Fukuyama, Japan
- Jörg Stülke, University of Göttingen, Germany Homepage
- Oscar Kuipers, University of Groningen, The Netherlands Homepage
Your additional remarks
References
Reviews
Sabine Brantl, Andreas Licht
Characterisation of Bacillus subtilis transcriptional regulators involved in metabolic processes.
Curr Protein Pept Sci: 2010, 11(4);274-91
[PubMed:20408793]
[WorldCat.org]
[DOI]
(I p)
Yasutaro Fujita
Carbon catabolite control of the metabolic network in Bacillus subtilis.
Biosci Biotechnol Biochem: 2009, 73(2);245-59
[PubMed:19202299]
[WorldCat.org]
[DOI]
(I p)
Boris Görke, Jörg Stülke
Carbon catabolite repression in bacteria: many ways to make the most out of nutrients.
Nat Rev Microbiol: 2008, 6(8);613-24
[PubMed:18628769]
[WorldCat.org]
[DOI]
(I p)
Josef Deutscher
The mechanisms of carbon catabolite repression in bacteria.
Curr Opin Microbiol: 2008, 11(2);87-93
[PubMed:18359269]
[WorldCat.org]
[DOI]
(P p)
Jessica B Warner, Juke S Lolkema
CcpA-dependent carbon catabolite repression in bacteria.
Microbiol Mol Biol Rev: 2003, 67(4);475-90
[PubMed:14665673]
[WorldCat.org]
[DOI]
(P p)
T M Henkin
The role of CcpA transcriptional regulator in carbon metabolism in Bacillus subtilis.
FEMS Microbiol Lett: 1996, 135(1);9-15
[PubMed:8598282]
[WorldCat.org]
[DOI]
(P p)
General and physiological studies
Global analyses (proteome, transcriptome)
Andrzej T Lulko, Girbe Buist, Jan Kok, Oscar P Kuipers
Transcriptome analysis of temporal regulation of carbon metabolism by CcpA in Bacillus subtilis reveals additional target genes.
J Mol Microbiol Biotechnol: 2007, 12(1-2);82-95
[PubMed:17183215]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
M S Moreno, B L Schneider, R R Maile, W Weyler, M H Saier
Catabolite repression mediated by the CcpA protein in Bacillus subtilis: novel modes of regulation revealed by whole-genome analyses.
Mol Microbiol: 2001, 39(5);1366-81
[PubMed:11251851]
[WorldCat.org]
[DOI]
(P p)
K Yoshida, K Kobayashi, Y Miwa, C M Kang, M Matsunaga, H Yamaguchi, S Tojo, M Yamamoto, R Nishi, N Ogasawara, T Nakayama, Y Fujita
Combined transcriptome and proteome analysis as a powerful approach to study genes under glucose repression in Bacillus subtilis.
Nucleic Acids Res: 2001, 29(3);683-92
[PubMed:11160890]
[WorldCat.org]
[DOI]
(I p)
S Tobisch, D Zühlke, J Bernhardt, J Stülke, M Hecker
Role of CcpA in regulation of the central pathways of carbon catabolism in Bacillus subtilis.
J Bacteriol: 1999, 181(22);6996-7004
[PubMed:10559165]
[WorldCat.org]
[DOI]
(P p)
Repression of target genes by CcpA
Positive regulation of gene expression by CcpA
Robert P Shivers, Abraham L Sonenshein
Bacillus subtilis ilvB operon: an intersection of global regulons.
Mol Microbiol: 2005, 56(6);1549-59
[PubMed:15916605]
[WorldCat.org]
[DOI]
(P p)
Holger Ludwig, Christoph Meinken, Anastasija Matin, Jörg Stülke
Insufficient expression of the ilv-leu operon encoding enzymes of branched-chain amino acid biosynthesis limits growth of a Bacillus subtilis ccpA mutant.
J Bacteriol: 2002, 184(18);5174-8
[PubMed:12193635]
[WorldCat.org]
[DOI]
(P p)
A J Turinsky, T R Moir-Blais, F J Grundy, T M Henkin
Bacillus subtilis ccpA gene mutants specifically defective in activation of acetoin biosynthesis.
J Bacteriol: 2000, 182(19);5611-4
[PubMed:10986270]
[WorldCat.org]
[DOI]
(P p)
E Presecan-Siedel, A Galinier, R Longin, J Deutscher, A Danchin, P Glaser, I Martin-Verstraete
Catabolite regulation of the pta gene as part of carbon flow pathways in Bacillus subtilis.
J Bacteriol: 1999, 181(22);6889-97
[PubMed:10559153]
[WorldCat.org]
[DOI]
(P p)
A J Turinsky, F J Grundy, J H Kim, G H Chambliss, T M Henkin
Transcriptional activation of the Bacillus subtilis ackA gene requires sequences upstream of the promoter.
J Bacteriol: 1998, 180(22);5961-7
[PubMed:9811655]
[WorldCat.org]
[DOI]
(P p)
F J Grundy, D A Waters, S H Allen, T M Henkin
Regulation of the Bacillus subtilis acetate kinase gene by CcpA.
J Bacteriol: 1993, 175(22);7348-55
[PubMed:8226682]
[WorldCat.org]
[DOI]
(P p)
Control of CcpA activity
CcpA-DNA interaction
Functional analysis of CcpA
Structural analyses