SrfAD
- Description: surfactin synthetase / competence
| Gene name | srfAD |
| Synonyms | comL |
| Essential | no |
| Product | surfactin synthetase / competence |
| Function | antibiotic synthesis |
| Gene expression levels in SubtiExpress: srfAD | |
| MW, pI | 27 kDa, 5.22 |
| Gene length, protein length | 726 bp, 242 aa |
| Immediate neighbours | srfAC, ycxA |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 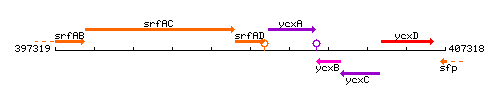
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed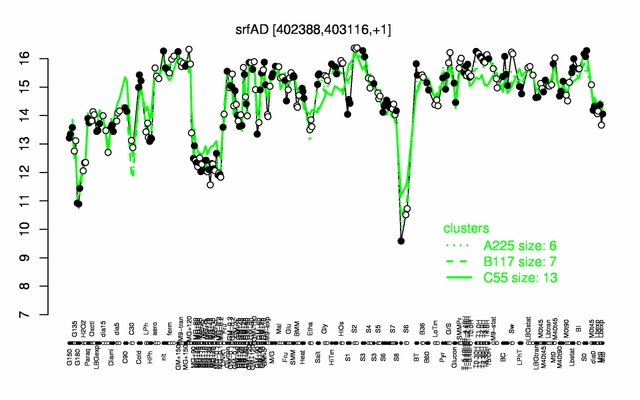
| |
Contents
Categories containing this gene/protein
miscellaneous metabolic pathways, biosynthesis of antibacterial compounds
This gene is a member of the following regulons
Abh regulon, CodY regulon, ComA regulon, PerR regulon, Spx regulon
The gene
Basic information
- Locus tag: BSU03520
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: thioesterase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: Q08788
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Additional publications: PubMed
Alexander Koglin, Frank Löhr, Frank Bernhard, Vladimir V Rogov, Dominique P Frueh, Eric R Strieter, Mohammad R Mofid, Peter Güntert, Gerhard Wagner, Christopher T Walsh, Mohamed A Marahiel, Volker Dötsch
Structural basis for the selectivity of the external thioesterase of the surfactin synthetase.
Nature: 2008, 454(7206);907-11
[PubMed:18704089]
[WorldCat.org]
[DOI]
(I p)
Mitsuo Ogura, Yasutaro Fujita
Bacillus subtilis rapD, a direct target of transcription repression by RghR, negatively regulates srfA expression.
FEMS Microbiol Lett: 2007, 268(1);73-80
[PubMed:17227471]
[WorldCat.org]
[DOI]
(P p)
Paul D Straight, Michael A Fischbach, Christopher T Walsh, David Z Rudner, Roberto Kolter
A singular enzymatic megacomplex from Bacillus subtilis.
Proc Natl Acad Sci U S A: 2007, 104(1);305-10
[PubMed:17190806]
[WorldCat.org]
[DOI]
(P p)
Kentaro Hayashi, Taku Ohsawa, Kazuo Kobayashi, Naotake Ogasawara, Mitsuo Ogura
The H2O2 stress-responsive regulator PerR positively regulates srfA expression in Bacillus subtilis.
J Bacteriol: 2005, 187(19);6659-67
[PubMed:16166527]
[WorldCat.org]
[DOI]
(P p)
Natalia Comella, Alan D Grossman
Conservation of genes and processes controlled by the quorum response in bacteria: characterization of genes controlled by the quorum-sensing transcription factor ComA in Bacillus subtilis.
Mol Microbiol: 2005, 57(4);1159-74
[PubMed:16091051]
[WorldCat.org]
[DOI]
(P p)
Shunji Nakano, Michiko M Nakano, Ying Zhang, Montira Leelakriangsak, Peter Zuber
A regulatory protein that interferes with activator-stimulated transcription in bacteria.
Proc Natl Acad Sci U S A: 2003, 100(7);4233-8
[PubMed:12642660]
[WorldCat.org]
[DOI]
(P p)
P Serror, A L Sonenshein
CodY is required for nutritional repression of Bacillus subtilis genetic competence.
J Bacteriol: 1996, 178(20);5910-5
[PubMed:8830686]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, L A Xia, P Zuber
Transcription initiation region of the srfA operon, which is controlled by the comP-comA signal transduction system in Bacillus subtilis.
J Bacteriol: 1991, 173(17);5487-93
[PubMed:1715856]
[WorldCat.org]
[DOI]
(P p)