YjmD
- Description: galactitol-1-phosphate dehydrogenase
| Gene name | yjmD |
| Synonyms | |
| Essential | no |
| Product | galactitol-1-phosphate dehydrogenase |
| Function | utilization of dulcitol |
| Gene expression levels in SubtiExpress: yjmD | |
| MW, pI | 36 kDa, 6.115 |
| Gene length, protein length | 1017 bp, 339 aa |
| Immediate neighbours | yjmC, uxuA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 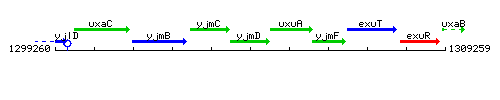
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed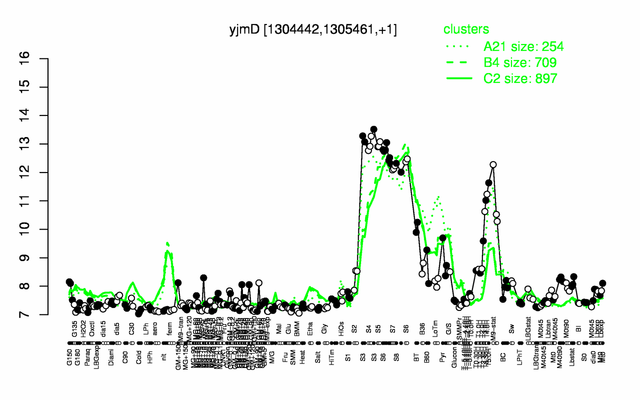
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, sporulation proteins
This gene is a member of the following regulons
CcpA regulon, ExuR regulon, SigE regulon
The gene
Basic information
- Locus tag: BSU12330
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: zinc-containing alcohol dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O35045
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Bogumiła C Marciniak, Monika Pabijaniak, Anne de Jong, Robert Dűhring, Gerald Seidel, Wolfgang Hillen, Oscar P Kuipers
High- and low-affinity cre boxes for CcpA binding in Bacillus subtilis revealed by genome-wide analysis.
BMC Genomics: 2012, 13;401
[PubMed:22900538]
[WorldCat.org]
[DOI]
(I e)
You-Kwan Oh, Bernhard O Palsson, Sung M Park, Christophe H Schilling, Radhakrishnan Mahadevan
Genome-scale reconstruction of metabolic network in Bacillus subtilis based on high-throughput phenotyping and gene essentiality data.
J Biol Chem: 2007, 282(39);28791-28799
[PubMed:17573341]
[WorldCat.org]
[DOI]
(P p)
Y Miwa, A Nakata, A Ogiwara, M Yamamoto, Y Fujita
Evaluation and characterization of catabolite-responsive elements (cre) of Bacillus subtilis.
Nucleic Acids Res: 2000, 28(5);1206-10
[PubMed:10666464]
[WorldCat.org]
[DOI]
(I p)
K R Mekjian, E M Bryan, B W Beall, C P Moran
Regulation of hexuronate utilization in Bacillus subtilis.
J Bacteriol: 1999, 181(2);426-33
[PubMed:9882655]
[WorldCat.org]
[DOI]
(P p)
Carlo Rivolta, Blazenka Soldo, Vladimir Lazarevic, Bernard Joris, Catherine Mauël, Dimitri Karamat
A 35.7 kb DNA fragment from the Bacillus subtilis chromosome containing a putative 12.3 kb operon involved in hexuronate catabolism and a perfectly symmetrical hypothetical catabolite-responsive element.
Microbiology (Reading): 1998, 144 ( Pt 4);877-884
[PubMed:9579062]
[WorldCat.org]
[DOI]
(P p)