Difference between revisions of "PckA"
| Line 33: | Line 33: | ||
__TOC__ | __TOC__ | ||
| − | <br/><br/> | + | <br/><br/><br/><br/><br/><br/> |
Revision as of 15:09, 22 June 2009
- Description: phosphoenolpyruvate carboxykinase
| Gene name | pckA |
| Synonyms | ppc |
| Essential | no |
| Product | phosphoenolpyruvate carboxykinase |
| Function | synthesis of phosphoenolpyruvate |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 58,1 kDa, 5.12 |
| Gene length, protein length | 1581 bp, 527 amino acids |
| Immediate neighbours | metK, ytmB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 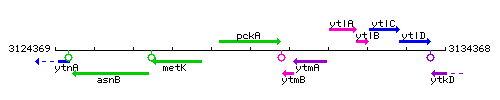
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU30560
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + oxaloacetate = ADP + phosphoenolpyruvate + CO2 (according to Swiss-Prot) ATP + oxaloacetate = ADP + phosphoenolpyruvate + CO(2)
- Protein family: phosphoenolpyruvate carboxykinase [ATP] family (according to Swiss-Prot) phosphoenolpyruvate carboxykinase [ATP] family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Nucleotide binding Domain (233–240)
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot), cytoplasm
Database entries
- Structure: 2PXZ (E.coli)
- Swiss prot entry: P54418
- KEGG entry: [3]
- E.C. number: 4.1.1.49
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism: transcription repression
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Your additional remarks
References