Difference between revisions of "IlvC"
| Line 122: | Line 122: | ||
=References= | =References= | ||
| − | <pubmed>15547269,12618455,12107147, </pubmed> | + | <pubmed>15060025,12193635,19258532,8289305,18641142,15547269,12618455,,15547269,12618455,12107147, </pubmed> |
# Mäder et al. (2002) Transcriptome and Proteome Analysis of ''Bacillus subtilis'' Gene Expression Modulated by Amino Acid Availability. ''J. Bacteriol'' '''184:''' 1844288-4295 [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | # Mäder et al. (2002) Transcriptome and Proteome Analysis of ''Bacillus subtilis'' Gene Expression Modulated by Amino Acid Availability. ''J. Bacteriol'' '''184:''' 1844288-4295 [http://www.ncbi.nlm.nih.gov/pubmed/12107147 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 23:34, 13 June 2009
- Description: ketol-acid reductoisomerase (2,3-dihydroxy-3-methylbutanoate, 2-acetolactate)
| Gene name | ilvC |
| Synonyms | |
| Essential | no |
| Product | ketol-acid reductoisomerase (2,3-dihydroxy-3-methylbutanoate, 2-acetolactate) |
| Function | biosynthesis of branched-chain amino acids |
| MW, pI | 37 kDa, 5.37 |
| Gene length, protein length | 1026 bp, 342 aa |
| Immediate neighbours | leuA, ilvH |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 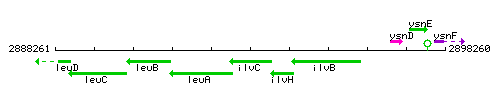
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU28290
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (R)-2,3-dihydroxy-3-methylbutanoate + NADP+ = (S)-2-hydroxy-2-methyl-3-oxobutanoate + NADPH (according to Swiss-Prot)
- Protein family: ketol-acid reductoisomerase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry: P37253
- KEGG entry: [3]
- E.C. number: 1.1.1.86
Additional information
Expression and regulation
- Regulation: repressed by casamino acids PubMed , expressed in the absence of branched-chain amino acids (BCAA), expression is stimulated in the presence of glucose PubMed, repressed by CodY PubMed
- Regulatory mechanism: CodY: transcription repression PubMed, glucose regulation: CcpA PubMed, repression by BCAA: tRNA-controlled RNA switch (T-box) that mediates termination/antitermination
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References