Difference between revisions of "Icd"
Cadu Cunha (talk | contribs) (→Additional information) |
Cadu Cunha (talk | contribs) (→Database entries) |
||
| Line 85: | Line 85: | ||
* '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU29130] | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU29130] | ||
| − | * '''E.C. number:''' [http://www.expasy.org/enzyme/1.1.1.42 1.1.1.42] | + | * '''E.C. number:''' [http://www.expasy.org/enzyme/1.1.1.42 1.1.1.42] |
=== Additional information=== | === Additional information=== | ||
Revision as of 15:22, 10 June 2009
- Description: isocitrate dehydrogenase
| Gene name | icd |
| Synonyms | citC |
| Essential | no |
| Product | isocitrate dehydrogenase |
| Function | TCA cycle |
| MW, pI | 46 kDa, 4.833 |
| Gene length, protein length | 1269 bp, 423 aa |
| Immediate neighbours | citZ, mdh |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 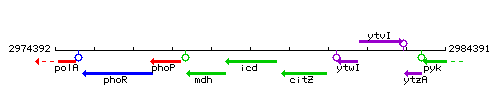
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU29130
Phenotypes of a mutant
reduced ability to sporulate PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Isocitrate + NADP+ = 2-oxoglutarate + CO2 + NADPH (according to Swiss-Prot)
- Protein family: isocitrate and isopropylmalate dehydrogenases family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Reversible Michaelis-Menten PubMed
- Domains:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: attached to the membrane PubMed
Database entries
- Structure: 1HQS
- Swiss prot entry: P39126
- KEGG entry: [3]
- E.C. number: 1.1.1.42
Additional information
This enzyme requires NADP+ exclusively. No activity was seen on the presence on NAD+ PubMed
Expression and regulation
- Sigma factor: SigA
- Regulation: repressed by glucose (2.2-fold) (CcpA) PubMed, citZ: catabolite repression (CcpA), repression by glucose + glutamate (CcpC)
icd: constitutive
- Regulatory mechanism: CcpA: transcription repression
- Additional information:
Biological materials
- Mutant: GP666 (spc), GP672 (erm), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Linc Sonenshein lab
Labs working on this gene/protein
[Linc Sonenshein|Linc Sonenshein]], Tufts University, Boston, MA, USA Homepage
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149 PubMed
- Matsuno K, Blais T, Serio AW, Conway T, Henkin TM, Sonenshein AL (1999) Metabolic imbalance and sporulation in an isocitrate dehydrogenase mutant of Bacillus subtilis. J Bacteriol 181:3382-3391. PubMed
- Singh SK, Miller SP, Dean A, Banaszak LJ, LaPorte DC (2002) Bacillus subtilis isocitrate dehydrogenase. A substrate analogue for Escherichia coli isocitrate dehydrogenase kinase/ phosphatase. J Biol Chem 277: 7567-7573. PubMed
- Hahne et al. (2008) From complementarity to comprehensiveness - targeting the membrane proteome of growing Bacillus subtilis by divergent approaches. Proteomics 8: 4123-4136 PubMed
- Eymann et al. (2007) Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis. Proteomics 7: 3509-3526. PubMed
- Levine et al. (2006) Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes. Proteomics 6, 2157-2173. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed