Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' N-acetylglucosamine-6-phosphate deacetylase <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''nagA'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Essential''' || | + | |style="background:#ABCDEF;" align="center"| '''Essential''' || no |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || N-acetylglucosamine-6-phosphate deacetylase |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || N-acetylglucosamine utilization |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 42 kDa, 5.276 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1188 bp, 396 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[hprK]]'', ''[[nagB]]'' |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS: | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB15506]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:nagA_context.gif]] |
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| Line 35: | Line 35: | ||
=== Basic information === | === Basic information === | ||
| − | * '''Locus tag:''' | + | * '''Locus tag:''' BSU35010 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | |||
| − | |||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' | + | * '''DBTBS entry:''' no entry |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG12630] |
=== Additional information=== | === Additional information=== | ||
| Line 54: | Line 52: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' N-acetyl-D-glucosamine 6-phosphate + H<sub>2</sub>O = D-glucosamine 6-phosphate + acetate (according to Swiss-Prot) |
| − | * '''Protein family:''' | + | * '''Protein family:''' nagA family (according to Swiss-Prot) |
* '''Paralogous protein(s):''' | * '''Paralogous protein(s):''' | ||
| Line 78: | Line 76: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId= | + | * '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=2VHL 2VHL] |
| − | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/ | + | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O34450 O34450] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+ | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU35010] |
| − | * '''E.C. number:''' [http://www.expasy.org/enzyme/ | + | * '''E.C. number:''' [http://www.expasy.org/enzyme/3.5.1.25 3.5.1.25] |
=== Additional information=== | === Additional information=== | ||
| Line 92: | Line 90: | ||
* '''Operon:''' | * '''Operon:''' | ||
| − | * '''[[Sigma factor]]:''' | + | * '''[[Sigma factor]]:''' |
| − | * '''Regulation:''' | + | * '''Regulation:''' repressed by glucose (2.9-fold) ([[CcpA]]) [http://www.ncbi.nlm.nih.gov/pubmed/12850135 PubMed] |
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' [[CcpA]]: transcription repression |
* '''Additional information:''' | * '''Additional information:''' | ||
| Line 120: | Line 118: | ||
=References= | =References= | ||
| − | <pubmed> | + | <pubmed>12850135 6174502, </pubmed> |
| − | # | + | # Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in ''Bacillus subtilis'': regulation of the central metabolic pathways. ''Metab Eng.'' '''5:''' 133-149 [http://www.ncbi.nlm.nih.gov/pubmed/12850135 PubMed] |
| + | # Mobley HL, Doyle RJ, Streips UN, Langemeier SO. (1982) Transport and incorporation of N-acetyl-D-glucosamine in Bacillus subtilis. ''J Bacteriol. '' '''Apr;150(1):''' 8-15. [http://www.ncbi.nlm.nih.gov/sites/entrez/6174502 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 13:26, 8 June 2009
- Description: N-acetylglucosamine-6-phosphate deacetylase
| Gene name | nagA |
| Synonyms | |
| Essential | no |
| Product | N-acetylglucosamine-6-phosphate deacetylase |
| Function | N-acetylglucosamine utilization |
| MW, pI | 42 kDa, 5.276 |
| Gene length, protein length | 1188 bp, 396 aa |
| Immediate neighbours | hprK, nagB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 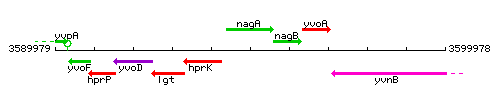
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU35010
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: N-acetyl-D-glucosamine 6-phosphate + H2O = D-glucosamine 6-phosphate + acetate (according to Swiss-Prot)
- Protein family: nagA family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 2VHL
- Swiss prot entry: O34450
- KEGG entry: [2]
- E.C. number: 3.5.1.25
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism: CcpA: transcription repression
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149 PubMed
- Mobley HL, Doyle RJ, Streips UN, Langemeier SO. (1982) Transport and incorporation of N-acetyl-D-glucosamine in Bacillus subtilis. J Bacteriol. Apr;150(1): 8-15. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed