Difference between revisions of "PrkC"
(→Expression and regulation) |
(→Expression and regulation) |
||
| Line 92: | Line 92: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[prpC]]-[[prkC]]-[[ | + | * '''Operon:''' ''[[prpC]]-[[prkC]]-[[cpgA]]'' [http://www.ncbi.nlm.nih.gov/pubmed/16025310?ordinalpos=9&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] |
* '''Sigma factor:''' | * '''Sigma factor:''' | ||
Revision as of 21:15, 22 May 2009
- Description: protein kinase C
| Gene name | prkC |
| Synonyms | yloP |
| Essential | no |
| Product | protein kinase |
| Function | germination in response to muropeptides |
| MW, pI | 71 kDa, 4.833 |
| Gene length, protein length | 1944 bp, 648 aa |
| Immediate neighbours | prpC, cpgA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 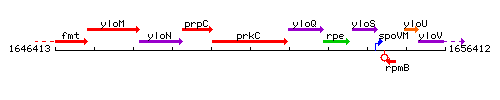
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: protein kinase domain (according to Swiss-Prot)
- Paralogous protein(s):
Proteins phosphorylated by PrkC
CpgA, EF-Tu, YezB PubMed, EF-G PubMed
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on Thr-290 PubMed
- Cofactor(s):
- Effectors of protein activity: activated by muropeptides PubMed
- Interactions:
- Localization: membrane (according to Swiss-Prot), membrane PubMed
Database entries
- Structure:
- Swiss prot entry: O34507
- KEGG entry: BSU15770
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector: pGP832 (N-terminal Strep-tag, purification from B. subtilis, for SPINE, in pGP380), available in Stülke lab
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Macek et al. (2007) The serine/ threonine/ tyrosine phosphoproteome of the model bacterium Bacillus subtilis. Mol. Cell. Proteomics 6: 697-707 PubMed
- Gaidenko TA, Kim TJ, Price CW: (2002) The PrpC serine-threonine phosphatase and PrkC kinase have opposing physiological roles in stationary-phase Bacillus subtilis cells. J Bacteriol, 184:6109-6114. PubMed
- Madec E, Laszkiewicz A, Iwanicki A, Obuchowski M, Séror S: (2002) Characterization of a membrane-linked Ser/Thr protein kinase in Bacillus subtilis, implicated in developmental processes. Mol Microbiol, 46:571-586. PubMed
- Madec E, Stensballe A, Kjellström S, Cladière L, Obuchowski M, Jensen ON, Séror S: (2003) Mass spectrometry and site-directed mutagenesis identify several autophosphorylated residues required for the activity of PrkC, a Ser/Thr kinase from Bacillus subtilis. J Mol Biol 330:459-472. PubMed
- Shah IM, Laaberki MH, Popham DL, Dworkin J: A eukaryotic-like Ser/Thr kinase signals bacteria to exit dormancy in response to peptidoglycan fragments. Cell 2008, 135:486-496. PubMed
- Absalon C, Obuchowski M, Madec E, Delattre D, Holland IB, Séror SJ (2009) CpgA, EF-Tu and the stressosome protein YezB are substrates of the Ser/Thr kinase/phosphatase couple, PrkC/PrpC, in Bacillus subtilis. Microbiology 155: 932-943. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed