Difference between revisions of "LeuD"
(→References) |
(→References) |
||
| Line 154: | Line 154: | ||
=References= | =References= | ||
| − | <pubmed>15060025,12193635,19258532,18697947,8289305,18641142,11948165,15547269,12618455,15547269,12618455,12107147, 25157083 20935095 24163341</pubmed> | + | <pubmed>15060025,12193635,19258532,18697947,8289305,18641142,11948165,15547269,12618455,15547269,12618455,12107147, 25157083 20935095 25755103 24163341</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 09:17, 30 July 2015
- Description: 3-isopropylmalate dehydratase (small subunit)
| Gene name | leuD |
| Synonyms | |
| Essential | no |
| Product | 3-isopropylmalate dehydratase (small subunit) |
| Function | biosynthesis of leucine |
| Gene expression levels in SubtiExpress: leuD | |
| Metabolic function and regulation of this protein in SubtiPathways: leuD | |
| MW, pI | 22 kDa, 4.582 |
| Gene length, protein length | 597 bp, 199 aa |
| Immediate neighbours | ysoA, leuC |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 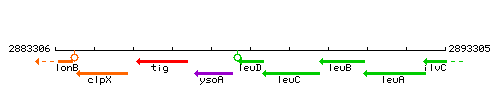
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed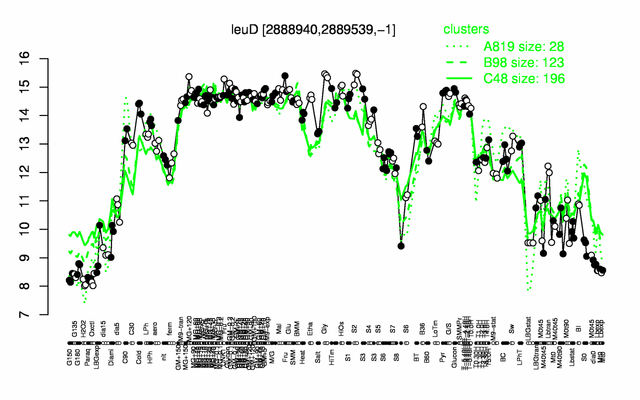
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
CcpA regulon, CodY regulon, FsrA regulon, T-box, TnrA regulon
The gene
Basic information
- Locus tag: BSU28250
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28250
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (2R,3S)-3-isopropylmalate = (2S)-2-isopropylmaleate + H2O (according to Swiss-Prot)
- Protein family: LeuD type 1 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Cofactors: contains an iron-sulfur cluster
- Effectors of protein activity:
Database entries
- BsubCyc: BSU28250
- Structure: 2PKP (from Methanocaldococcus jannaschii dsm 2661, 38% identity, 53% similarity)
- UniProt: P94568
- KEGG entry: [3]
- E.C. number: 4.2.1.33
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- for a complete overview on the regulation of the ilv operon, see Brinsmade et al.
- repressed by casamino acids PubMed
- expression is stimulated in the presence of glucose PubMed
- repressed in the absence of good nitrogen sources (glutamine or ammonium) (TnrA) PubMed
- repressed during growth in the presence of branched chain amino acids (CodY) PubMed
- less expressed under conditions of extreme iron limitation (FsrA) PubMed
- Regulatory mechanism:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 1251 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 5846 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 3609 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 2313 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References