Difference between revisions of "YidC2"
(→Original publications) |
|||
| Line 138: | Line 138: | ||
<pubmed>22688815 20800571,21194367,15802250</pubmed> | <pubmed>22688815 20800571,21194367,15802250</pubmed> | ||
== Original publications == | == Original publications == | ||
| − | <pubmed>17114254, 15995216, 11889108, 12586834 19717609 19779460 21204254 22383849 22864117 24443530 25313395 </pubmed> | + | <pubmed>17114254, 15995216, 11889108, 12586834 19717609 19779460 21204254 22383849 22864117 24443530 25313395 25359772 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 15:41, 2 November 2014
- Description: Sec-independent membrane protein translocase, required for development of genetic competence
| Gene name | yidC2 |
| Synonyms | yqjG, oxaAB |
| Essential | no |
| Product | Sec-independent membrane protein translocase |
| Function | membrane insertion of proteins and protein secretion |
| MW, pI | 30 kDa, 9.903 |
| Gene length, protein length | 825 bp, 275 aa |
| Immediate neighbours | mifM, yqjF |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 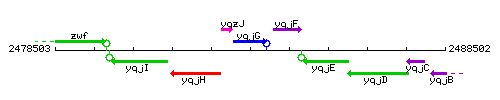
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed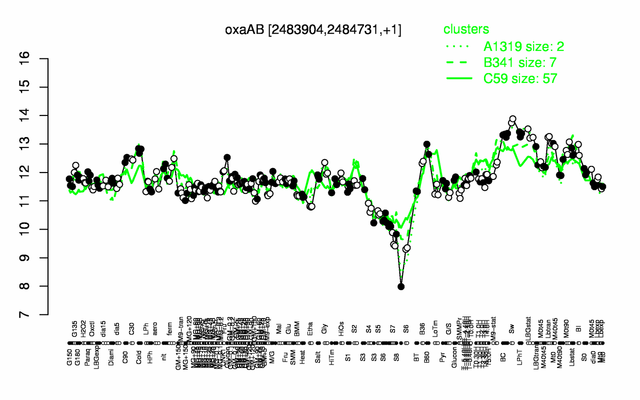
| |
Contents
Categories containing this gene/protein
protein secretion, genetic competence, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23890
Phenotypes of a mutant
Database entries
- BsubCyc: BSU23890
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: YidC/Oxa1/Alb3 family PubMed
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization:
- membrane PubMed
Database entries
- BsubCyc: BSU23890
- Structure:
- UniProt: P54544
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications