Difference between revisions of "YjbH"
(→Original Publications) |
|||
| Line 149: | Line 149: | ||
<pubmed> 23375660 23479438,19609260</pubmed> | <pubmed> 23375660 23479438,19609260</pubmed> | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>17293416, 19074380 17908206, 20525796 21378193,21947404 24417481</pubmed> | + | <pubmed>17293416, 19074380 17908206, 20525796 21378193,21947404 24417481 24942655 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:51, 20 June 2014
- Description: adaptor protein for ClpX-ClpP-catalyzed Spx degradation, confers resistance against nitrosating agents
| Gene name | yjbH |
| Synonyms | |
| Essential | no |
| Product | adaptor protein |
| Function | stimulation of Spx degradation |
| Gene expression levels in SubtiExpress: yjbH | |
| Interactions involving this protein in SubtInteract: YjbH | |
| MW, pI | 31 kDa, 5.206 |
| Gene length, protein length | 825 bp, 275 aa |
| Immediate neighbours | yizD, yjbI |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 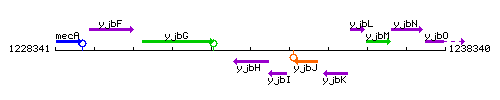
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed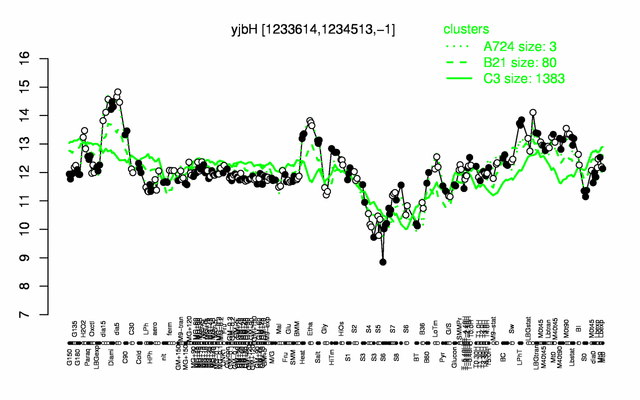
| |
Contents
Categories containing this gene/protein
proteolysis, resistance against other toxic compounds (nitric oxide, phenolic acids, flavonoids, oxalate)
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU11550
Phenotypes of a mutant
- increased thermotolerance due to increased stabiliy of Spx and thus increased expression of trxA PubMed
Database entries
- BsubCyc: BSU11550
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: adaptor protein for ClpX-ClpP-catalyzed Spx degradation PubMed
- Protein family: UPF0413 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization: cytosolic protein
Database entries
- BsubCyc: BSU11550
- Structure:
- UniProt: O31606
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Peter Zuber, Oregon Health and Science University, USA Homepage
Claes von Wachenfeldt, Lund University, Sweden Homepage
Your additional remarks
References
Reviews
Noël Molière, Kürşad Turgay
General and regulatory proteolysis in Bacillus subtilis.
Subcell Biochem: 2013, 66;73-103
[PubMed:23479438]
[WorldCat.org]
[DOI]
(P p)
Aurelia Battesti, Susan Gottesman
Roles of adaptor proteins in regulation of bacterial proteolysis.
Curr Opin Microbiol: 2013, 16(2);140-7
[PubMed:23375660]
[WorldCat.org]
[DOI]
(I p)
Janine Kirstein, Noël Molière, David A Dougan, Kürşad Turgay
Adapting the machine: adaptor proteins for Hsp100/Clp and AAA+ proteases.
Nat Rev Microbiol: 2009, 7(8);589-99
[PubMed:19609260]
[WorldCat.org]
[DOI]
(I p)
Original Publications