Difference between revisions of "RecU"
| Line 1: | Line 1: | ||
| − | * '''Description:''' DNA repair, homologous recombination and chromosome segregation; recognizes, distorts, and cleaves four-stranded recombination intermediates and modulates [[RecA]] activities, equivalent to ''E. coli'' RecU<br/><br/> | + | * '''Description:''' Holliday junction resolvase, DNA repair, homologous recombination and chromosome segregation; recognizes, distorts, and cleaves four-stranded recombination intermediates and modulates [[RecA]] activities, equivalent to ''E. coli'' RecU<br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || Holliday junction resolvase | |style="background:#ABCDEF;" align="center"| '''Product''' || Holliday junction resolvase | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || DNA repair | + | |style="background:#ABCDEF;" align="center"|'''Function''' || [[DNA repair/ recombination]] and [[chromosome segregation]] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU22310 recU] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU22310 recU] | ||
| Line 53: | Line 53: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | * the inactivations of ''[[recU]]'' and ''[[ruvA]]-[[ruvB]]'' are synthetically lethal {{PubMed|24770420}} | ||
=== Database entries === | === Database entries === | ||
| Line 62: | Line 63: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 146: | Line 144: | ||
<pubmed> 22933559 9442895</pubmed> | <pubmed> 22933559 9442895</pubmed> | ||
== Original publications == | == Original publications == | ||
| − | <pubmed>9642195,16024744,15317759,11810266,19730681 , 19422832 7814321 18179421 16154091 21600217 18684995 16020779 </pubmed> | + | <pubmed>9642195,16024744,15317759,11810266,19730681 , 19422832 7814321 18179421 16154091 21600217 18684995 16020779 24770420 </pubmed> |
==RecU in other organisms== | ==RecU in other organisms== | ||
<pubmed>23356868 </pubmed> | <pubmed>23356868 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:27, 5 May 2014
- Description: Holliday junction resolvase, DNA repair, homologous recombination and chromosome segregation; recognizes, distorts, and cleaves four-stranded recombination intermediates and modulates RecA activities, equivalent to E. coli RecU
| Gene name | recU |
| Synonyms | recG, prfA, yppB, jopB |
| Essential | no |
| Product | Holliday junction resolvase |
| Function | DNA repair/ recombination and chromosome segregation |
| Gene expression levels in SubtiExpress: recU | |
| Interactions involving this protein in SubtInteract: RecU | |
| MW, pI | 23 kDa, 9.511 |
| Gene length, protein length | 618 bp, 206 aa |
| Immediate neighbours | yppC, ponA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 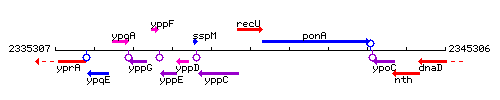
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed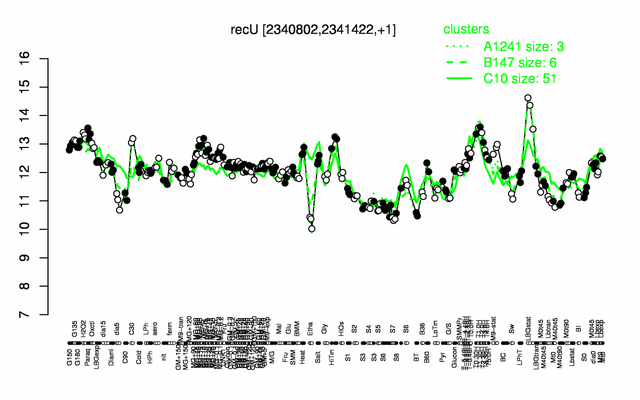
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, cell envelope stress proteins (controlled by SigM, V, W, X, Y)
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22310
Phenotypes of a mutant
Database entries
- BsubCyc: BSU22310
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- BsubCyc: BSU22310
- Structure: 1ZP7
- UniProt: P39792
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP891 (recU::cat), available in Jörg Stülke's lab
- 1A895 (no resistance), PubMed, available at BGSC
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
RecU in other organisms