Difference between revisions of "Apt"
| Line 127: | Line 127: | ||
** number of protein molecules per cell (minimal medium with glucose and ammonium): 872 {{PubMed|24696501}} | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 872 {{PubMed|24696501}} | ||
** number of protein molecules per cell (complex medium with amino acids, without glucose): 2750 {{PubMed|24696501}} | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 2750 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 1189 {{PubMed|21395229}} | ||
| + | |||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 1706 {{PubMed|21395229}} | ||
=Biological materials = | =Biological materials = | ||
| − | |||
* '''Mutant:''' | * '''Mutant:''' | ||
Revision as of 13:32, 17 April 2014
- Description: adenine phosphoribosyltransferase, universally conserved protein
| Gene name | apt |
| Synonyms | |
| Essential | no |
| Product | adenine phosphoribosyltransferase |
| Function | purine salvage and interconversion |
| Gene expression levels in SubtiExpress: apt | |
| Metabolic function and regulation of this protein in SubtiPathways: apt | |
| MW, pI | 18 kDa, 4.701 |
| Gene length, protein length | 510 bp, 170 aa |
| Immediate neighbours | relA, yrvE |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 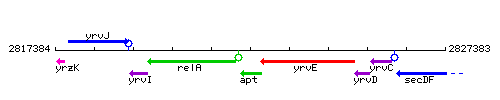
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed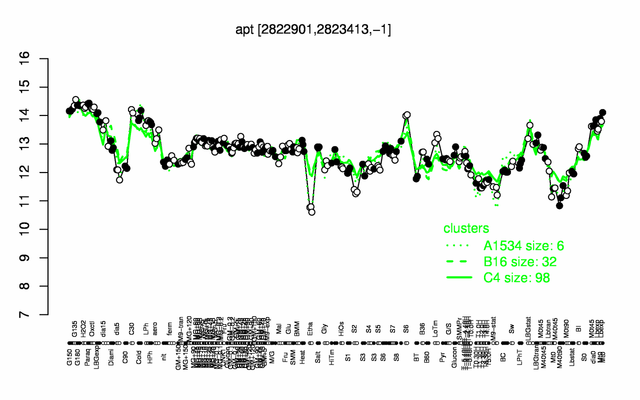
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of nucleotides, universally conserved proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU27610
Phenotypes of a mutant
Database entries
- BsubCyc: BSU27610
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: AMP + diphosphate = adenine + 5-phospho-alpha-D-ribose 1-diphosphate (according to Swiss-Prot)
- Protein family: purine/pyrimidine phosphoribosyltransferase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-85 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU27610
- Structure: 2DY0 (from Escherichia coli k12, 56% identity, 66% similarity)
- UniProt: O34443
- KEGG entry: [3]
- E.C. number: 2.4.2.7
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 872 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 2750 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 1189 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 1706 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References