Difference between revisions of "MetE"
| Line 129: | Line 129: | ||
** subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ** subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ||
** belongs to the 100 [[most abundant proteins]] {{PubMed|15378759}} | ** belongs to the 100 [[most abundant proteins]] {{PubMed|15378759}} | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 10685 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 1682 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 09:23, 17 April 2014
- Description: methionine synthase
| Gene name | metE |
| Synonyms | metC |
| Essential | no |
| Product | methionine synthase |
| Function | biosynthesis of methionine |
| Gene expression levels in SubtiExpress: metE | |
| Interactions involving this protein in SubtInteract: MetE | |
| Metabolic function and regulation of this protein in SubtiPathways: metE | |
| MW, pI | 86 kDa, 4.839 |
| Gene length, protein length | 2286 bp, 762 aa |
| Immediate neighbours | guaD, ispA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 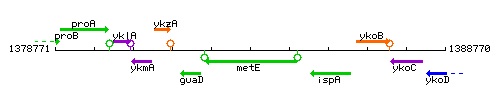
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed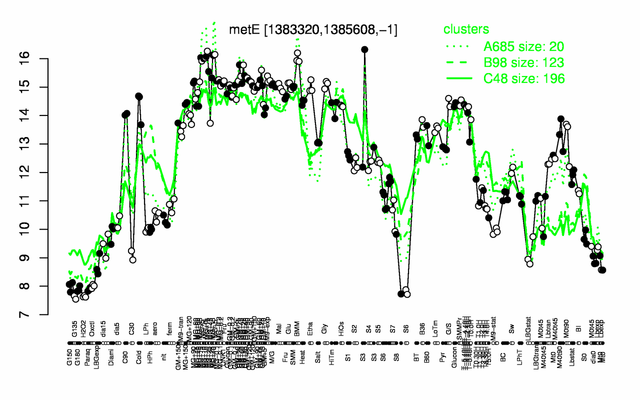
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13180
Phenotypes of a mutant
Database entries
- BsubCyc: BSU13180
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 5-methyltetrahydropteroyltri-L-glutamate + L-homocysteine = tetrahydropteroyltri-L-glutamate + L-methionine (according to Swiss-Prot)
- Protein family: vitamin-B12 independent methionine synthase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- phosphorylated on ser/ thr/ tyr PubMed, S-cysteinylation after diamide stress (C719) PubMed
- Cys719 and Cys730 are S-bacillithiolated by NaOCl stress in B. subtilis and other Bacillus species PubMed PubMed
- MetE is generally most strongly S-bacillithiolated by NaOCl stress in B. subtilis and other Bacillus species PubMed PubMed
- Effectors of protein activity:
Database entries
- BsubCyc: BSU13180
- UniProt: P80877
- KEGG entry: [2]
- E.C. number: 2.1.1.14
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon: metE
- Regulation:
- Regulatory mechanism: S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- Additional information:
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 10685 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 1682 PubMed
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Ahmed Gaballa, Bui Khanh Chi, Alexandra A Roberts, Dörte Becher, Chris J Hamilton, Haike Antelmann, John D Helmann
Redox regulation in Bacillus subtilis: The bacilliredoxins BrxA(YphP) and BrxB(YqiW) function in de-bacillithiolation of S-bacillithiolated OhrR and MetE.
Antioxid Redox Signal: 2014, 21(3);357-67
[PubMed:24313874]
[WorldCat.org]
[DOI]
(I p)
Allison Kriel, Shaun R Brinsmade, Jessica L Tse, Ashley K Tehranchi, Alycia N Bittner, Abraham L Sonenshein, Jue D Wang
GTP dysregulation in Bacillus subtilis cells lacking (p)ppGpp results in phenotypic amino acid auxotrophy and failure to adapt to nutrient downshift and regulate biosynthesis genes.
J Bacteriol: 2014, 196(1);189-201
[PubMed:24163341]
[WorldCat.org]
[DOI]
(I p)
Bui Khanh Chi, Alexandra A Roberts, Tran Thi Thanh Huyen, Katrin Bäsell, Dörte Becher, Dirk Albrecht, Chris J Hamilton, Haike Antelmann
S-bacillithiolation protects conserved and essential proteins against hypochlorite stress in firmicutes bacteria.
Antioxid Redox Signal: 2013, 18(11);1273-95
[PubMed:22938038]
[WorldCat.org]
[DOI]
(I p)
Bui Khanh Chi, Katrin Gronau, Ulrike Mäder, Bernd Hessling, Dörte Becher, Haike Antelmann
S-bacillithiolation protects against hypochlorite stress in Bacillus subtilis as revealed by transcriptomics and redox proteomics.
Mol Cell Proteomics: 2011, 10(11);M111.009506
[PubMed:21749987]
[WorldCat.org]
[DOI]
(I p)
Ana Gutiérrez-Preciado, Tina M Henkin, Frank J Grundy, Charles Yanofsky, Enrique Merino
Biochemical features and functional implications of the RNA-based T-box regulatory mechanism.
Microbiol Mol Biol Rev: 2009, 73(1);36-61
[PubMed:19258532]
[WorldCat.org]
[DOI]
(I p)
Jerneja Tomsic, Brooke A McDaniel, Frank J Grundy, Tina M Henkin
Natural variability in S-adenosylmethionine (SAM)-dependent riboswitches: S-box elements in bacillus subtilis exhibit differential sensitivity to SAM In vivo and in vitro.
J Bacteriol: 2008, 190(3);823-33
[PubMed:18039762]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Falko Hochgräfe, Jörg Mostertz, Dierk-Christoph Pöther, Dörte Becher, John D Helmann, Michael Hecker
S-cysteinylation is a general mechanism for thiol protection of Bacillus subtilis proteins after oxidative stress.
J Biol Chem: 2007, 282(36);25981-5
[PubMed:17611193]
[WorldCat.org]
[DOI]
(P p)
Falko Hochgräfe, Jörg Mostertz, Dirk Albrecht, Michael Hecker
Fluorescence thiol modification assay: oxidatively modified proteins in Bacillus subtilis.
Mol Microbiol: 2005, 58(2);409-25
[PubMed:16194229]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
F J Grundy, T M Henkin
The S box regulon: a new global transcription termination control system for methionine and cysteine biosynthesis genes in gram-positive bacteria.
Mol Microbiol: 1998, 30(4);737-49
[PubMed:10094622]
[WorldCat.org]
[DOI]
(P p)