Difference between revisions of "Rnc"
| Line 59: | Line 59: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU15930&redirect=T BSU15930] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/rnc-smc-ftsY.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/rnc-smc-ftsY.html] | ||
| Line 98: | Line 99: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU15930&redirect=T BSU15930] | ||
* '''Structure:''' | * '''Structure:''' | ||
Revision as of 13:42, 2 April 2014
- Description: RNase III; required for the degradation of sense/antisense transcripts in Gram positive, low GC bacteria
| Gene name | rnc |
| Synonyms | rncS |
| Essential | yes |
| Product | endoribonuclease III |
| Function | cleaves both 5'- and 3'-sites of the small cytoplasmic RNA precursor (scr) |
| Gene expression levels in SubtiExpress: rnc | |
| Metabolic function and regulation of this protein in SubtiPathways: Protein secretion | |
| MW, pI | 28 kDa, 8.076 |
| Gene length, protein length | 747 bp, 249 aa |
| Immediate neighbours | acpA, smc |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 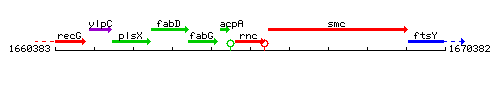
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed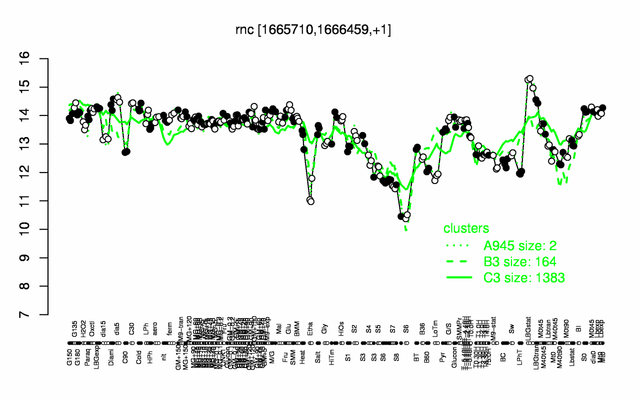
| |
Contents
Categories containing this gene/protein
Rnases, translation, protein secretion, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15930
Phenotypes of a mutant
- essential PubMed
- the rnc gene can be deleted in strains cured of the Skin element and SPß prophage (or upon deletion of the toxin genes txpA and yonT) PubMed
Database entries
- BsubCyc: BSU15930
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Endonucleolytic cleavage of double-stranded RNA to 5'-phosphomonoester (according to Swiss-Prot)
- cleaves both 5'- and 3'-sites of the small cytoplasmic RNA precursor (scr)
- degradation of txpA mRNA-RatA RNA duplex (encoded on the Skin element) PubMed
- degradation of yonT mRNA-as-yonT RNA duplex (encoded on the SP-beta prophage) PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU15930
- Structure:
- UniProt: P51833
- KEGG entry: [3]
- E.C. number: 3.1.26.3
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
David Bechhofer, Mount Sinai School, New York, USA Homepage
Ciaran Condon, IBPC, Paris, France Homepage
Your additional remarks
References
Reviews
Original publications
Durand S, Gilet L, Condon C The essential function of B. subtilis RNase III is to silence foreign toxin genes. PLoS Genet. 2012 8(12): e1003181. PubMed:23300471
Sylvain Durand, Laetitia Gilet, Philippe Bessières, Pierre Nicolas, Ciarán Condon
Three essential ribonucleases-RNase Y, J1, and III-control the abundance of a majority of Bacillus subtilis mRNAs.
PLoS Genet: 2012, 8(3);e1002520
[PubMed:22412379]
[WorldCat.org]
[DOI]
(I p)
Iñigo Lasa, Alejandro Toledo-Arana, Alexander Dobin, Maite Villanueva, Igor Ruiz de los Mozos, Marta Vergara-Irigaray, Víctor Segura, Delphine Fagegaltier, José R Penadés, Jaione Valle, Cristina Solano, Thomas R Gingeras
Genome-wide antisense transcription drives mRNA processing in bacteria.
Proc Natl Acad Sci U S A: 2011, 108(50);20172-7
[PubMed:22123973]
[WorldCat.org]
[DOI]
(I p)
Shiyi Yao, Joshua B Blaustein, David H Bechhofer
Processing of Bacillus subtilis small cytoplasmic RNA: evidence for an additional endonuclease cleavage site.
Nucleic Acids Res: 2007, 35(13);4464-73
[PubMed:17576666]
[WorldCat.org]
[DOI]
(I p)
M A Herskovitz, D H Bechhofer
Endoribonuclease RNase III is essential in Bacillus subtilis.
Mol Microbiol: 2000, 38(5);1027-33
[PubMed:11123676]
[WorldCat.org]
[DOI]
(P p)
Hiroshi Kakeshita, Akihiro Oguro, Reiko Amikura, Kouji Nakamura, Kunio Yamane
Expression of the ftsY gene, encoding a homologue of the alpha subunit of mammalian signal recognition particle receptor, is controlled by different promoters in vegetative and sporulating cells of Bacillus subtilis.
Microbiology (Reading): 2000, 146 ( Pt 10);2595-2603
[PubMed:11021934]
[WorldCat.org]
[DOI]
(P p)
A Oguro, H Kakeshita, K Nakamura, K Yamane, W Wang, D H Bechhofer
Bacillus subtilis RNase III cleaves both 5'- and 3'-sites of the small cytoplasmic RNA precursor.
J Biol Chem: 1998, 273(31);19542-7
[PubMed:9677377]
[WorldCat.org]
[DOI]
(P p)
W Wang, D H Bechhofer
Bacillus subtilis RNase III gene: cloning, function of the gene in Escherichia coli, and construction of Bacillus subtilis strains with altered rnc loci.
J Bacteriol: 1997, 179(23);7379-85
[PubMed:9393702]
[WorldCat.org]
[DOI]
(P p)
S Mitra, D H Bechhofer
Substrate specificity of an RNase III-like activity from Bacillus subtilis.
J Biol Chem: 1994, 269(50);31450-6
[PubMed:7527387]
[WorldCat.org]
(P p)
A T Panganiban, H R Whiteley
Bacillus subtilis RNAase III cleavage sites in phage SP82 early mRNA.
Cell: 1983, 33(3);907-13
[PubMed:6409421]
[WorldCat.org]
[DOI]
(P p)