Difference between revisions of "RsbT"
| Line 56: | Line 56: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU04690&redirect=T BSU04690] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/rsbRSTUVW-sigB-rsbX.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/rsbRSTUVW-sigB-rsbX.html] | ||
| Line 101: | Line 102: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU04690&redirect=T BSU04690] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=3VY9 3VY9] (complete stressosome) | * '''Structure:''' [http://www.rcsb.org/pdb/explore.do?structureId=3VY9 3VY9] (complete stressosome) | ||
Revision as of 13:02, 2 April 2014
- Description: PP2C activator, protein serine kinase, phosphorylates RsbS and RsbR, part of the stressosome
| Gene name | rsbT |
| Synonyms | ycxT |
| Essential | no |
| Product | PP2C activator, protein serine kinase |
| Function | control of SigB activity |
| Gene expression levels in SubtiExpress: rsbT | |
| Interactions involving this protein in SubtInteract: RsbT | |
| Metabolic function and regulation of this protein in SubtiPathways: rsbT | |
| MW, pI | 14 kDa, 6.587 |
| Gene length, protein length | 399 bp, 133 aa |
| Immediate neighbours | rsbS, rsbU |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 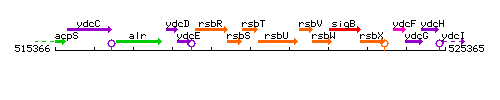
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed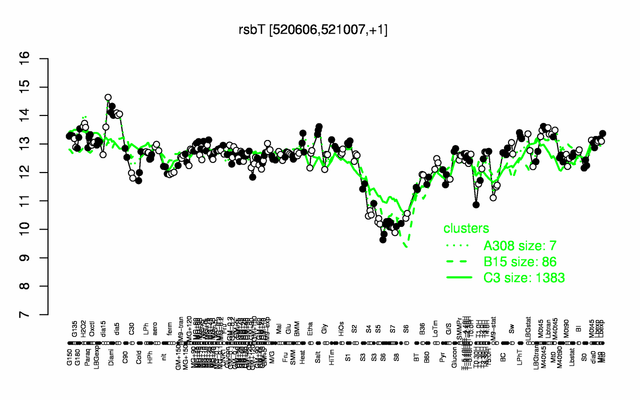
| |
Contents
Categories containing this gene/protein
protein modification, sigma factors and their control
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU04690
Phenotypes of a mutant
Database entries
- BsubCyc: BSU04690
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- component of the stressosome
Database entries
- BsubCyc: BSU04690
- Structure: 3VY9 (complete stressosome)
- UniProt: P42411
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation: constitutively expressed PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Bill Haldenwang, San Antonio, USA
- Chet Price, Davis, USA homepage
- Rick Lewis, Newcastle, UK homepage
Your additional remarks
References
Reviews
Original Articles
Tatiana A Gaidenko, Chester W Price
Genetic evidence for a phosphorylation-independent signal transduction mechanism within the Bacillus subtilis stressosome.
PLoS One: 2014, 9(3);e90741
[PubMed:24599254]
[WorldCat.org]
[DOI]
(I e)
Marcel Jurk, Philipp Schramm, Peter Schmieder
The blue-light receptor YtvA from Bacillus subtilis is permanently incorporated into the stressosome independent of the illumination state.
Biochem Biophys Res Commun: 2013, 432(3);499-503
[PubMed:23416074]
[WorldCat.org]
[DOI]
(I p)
Jonathan W Young, James C W Locke, Michael B Elowitz
Rate of environmental change determines stress response specificity.
Proc Natl Acad Sci U S A: 2013, 110(10);4140-5
[PubMed:23407164]
[WorldCat.org]
[DOI]
(I p)
Ulf W Liebal, Thomas Millat, Jon Marles-Wright, Richard J Lewis, Olaf Wolkenhauer
Simulations of stressosome activation emphasize allosteric interactions between RsbR and RsbT.
BMC Syst Biol: 2013, 7;3
[PubMed:23320651]
[WorldCat.org]
[DOI]
(I e)
James C W Locke, Jonathan W Young, Michelle Fontes, María Jesús Hernández Jiménez, Michael B Elowitz
Stochastic pulse regulation in bacterial stress response.
Science: 2011, 334(6054);366-9
[PubMed:21979936]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Stephan Schulz, Katrin Gronau, Dörte Becher, Michael Hecker, Chester W Price
In vivo phosphorylation patterns of key stressosome proteins define a second feedback loop that limits activation of Bacillus subtilis σB.
Mol Microbiol: 2011, 80(3);798-810
[PubMed:21362065]
[WorldCat.org]
[DOI]
(I p)
Adam Reeves, Luis Martinez, William Haldenwang
Expression of, and in vivo stressosome formation by, single members of the RsbR protein family in Bacillus subtilis.
Microbiology (Reading): 2010, 156(Pt 4);990-998
[PubMed:20019076]
[WorldCat.org]
[DOI]
(I p)
Jon Marles-Wright, Tim Grant, Olivier Delumeau, Gijs van Duinen, Susan J Firbank, Peter J Lewis, James W Murray, Joseph A Newman, Maureen B Quin, Paul R Race, Alexis Rohou, Willem Tichelaar, Marin van Heel, Richard J Lewis
Molecular architecture of the "stressosome," a signal integration and transduction hub.
Science: 2008, 322(5898);92-6
[PubMed:18832644]
[WorldCat.org]
[DOI]
(I p)
Steven W Hardwick, Jan Pané-Farré, Olivier Delumeau, Jon Marles-Wright, James W Murray, Michael Hecker, Richard J Lewis
Structural and functional characterization of partner switching regulating the environmental stress response in Bacillus subtilis.
J Biol Chem: 2007, 282(15);11562-72
[PubMed:17303566]
[WorldCat.org]
[DOI]
(P p)
Shuyu Zhang, Adam Reeves, Robyn L Woodbury, W G Haldenwang
Coexpression patterns of sigma(B) regulators in Bacillus subtilis affect sigma(B) inducibility.
J Bacteriol: 2005, 187(24);8520-5
[PubMed:16321960]
[WorldCat.org]
[DOI]
(P p)
Shrin Kuo, Shuyu Zhang, Robyn L Woodbury, W G Haldenwang
Associations between Bacillus subtilis sigmaB regulators in cell extracts.
Microbiology (Reading): 2004, 150(Pt 12);4125-36
[PubMed:15583165]
[WorldCat.org]
[DOI]
(P p)
Tae-Jong Kim, Tatiana A Gaidenko, Chester W Price
In vivo phosphorylation of partner switching regulators correlates with stress transmission in the environmental signaling pathway of Bacillus subtilis.
J Bacteriol: 2004, 186(18);6124-32
[PubMed:15342582]
[WorldCat.org]
[DOI]
(P p)
Tae-Jong Kim, Tatiana A Gaidenko, Chester W Price
A multicomponent protein complex mediates environmental stress signaling in Bacillus subtilis.
J Mol Biol: 2004, 341(1);135-50
[PubMed:15312768]
[WorldCat.org]
[DOI]
(P p)
Robyn L Woodbury, Tingqiu Luo, Lindsay Grant, W G Haldenwang
Mutational analysis of RsbT, an activator of the Bacillus subtilis stress response transcription factor, sigmaB.
J Bacteriol: 2004, 186(9);2789-97
[PubMed:15090521]
[WorldCat.org]
[DOI]
(P p)
Chien-Cheng Chen, Richard J Lewis, Robin Harris, Michael D Yudkin, Olivier Delumeau
A supramolecular complex in the environmental stress signalling pathway of Bacillus subtilis.
Mol Microbiol: 2003, 49(6);1657-69
[PubMed:12950928]
[WorldCat.org]
[DOI]
(P p)
Sujit Dutta, Richard J Lewis
Crystallization and preliminary crystallographic analysis of the kinase-recruitment domain of the PP2C-type phosphatase RsbU.
Acta Crystallogr D Biol Crystallogr: 2003, 59(Pt 1);191-3
[PubMed:12499568]
[WorldCat.org]
[DOI]
(P p)
S Zhang, J M Scott, W G Haldenwang
Loss of ribosomal protein L11 blocks stress activation of the Bacillus subtilis transcription factor sigma(B).
J Bacteriol: 2001, 183(7);2316-21
[PubMed:11244072]
[WorldCat.org]
[DOI]
(P p)
J M Scott, J Ju, T Mitchell, W G Haldenwang
The Bacillus subtilis GTP binding protein obg and regulators of the sigma(B) stress response transcription factor cofractionate with ribosomes.
J Bacteriol: 2000, 182(10);2771-7
[PubMed:10781545]
[WorldCat.org]
[DOI]
(P p)
T A Gaidenko, X Yang, Y M Lee, C W Price
Threonine phosphorylation of modulator protein RsbR governs its ability to regulate a serine kinase in the environmental stress signaling pathway of Bacillus subtilis.
J Mol Biol: 1999, 288(1);29-39
[PubMed:10329124]
[WorldCat.org]
[DOI]
(P p)
C M Kang, K Vijay, C W Price
Serine kinase activity of a Bacillus subtilis switch protein is required to transduce environmental stress signals but not to activate its target PP2C phosphatase.
Mol Microbiol: 1998, 30(1);189-96
[PubMed:9786195]
[WorldCat.org]
[DOI]
(P p)
N Smirnova, J Scott, U Voelker, W G Haldenwang
Isolation and characterization of Bacillus subtilis sigB operon mutations that suppress the loss of the negative regulator RsbX.
J Bacteriol: 1998, 180(14);3671-80
[PubMed:9658013]
[WorldCat.org]
[DOI]
(P p)
U Voelker, A Voelker, W G Haldenwang
The yeast two-hybrid system detects interactions between Bacillus subtilis sigmaB regulators.
J Bacteriol: 1996, 178(23);7020-3
[PubMed:8955331]
[WorldCat.org]
[DOI]
(P p)
X Yang, C M Kang, M S Brody, C W Price
Opposing pairs of serine protein kinases and phosphatases transmit signals of environmental stress to activate a bacterial transcription factor.
Genes Dev: 1996, 10(18);2265-75
[PubMed:8824586]
[WorldCat.org]
[DOI]
(P p)
U Voelker, A Voelker, W G Haldenwang
Reactivation of the Bacillus subtilis anti-sigma B antagonist, RsbV, by stress- or starvation-induced phosphatase activities.
J Bacteriol: 1996, 178(18);5456-63
[PubMed:8808936]
[WorldCat.org]
[DOI]
(P p)
C M Kang, M S Brody, S Akbar, X Yang, C W Price
Homologous pairs of regulatory proteins control activity of Bacillus subtilis transcription factor sigma(b) in response to environmental stress.
J Bacteriol: 1996, 178(13);3846-53
[PubMed:8682789]
[WorldCat.org]
[DOI]
(P p)
A Dufour, U Voelker, A Voelker, W G Haldenwang
Relative levels and fractionation properties of Bacillus subtilis σ(B) and its regulators during balanced growth and stress.
J Bacteriol: 1996, 178(13);3701-9 sigma
[PubMed:8682769]
[WorldCat.org]
[DOI]
(P p)
A A Wise, C W Price
Four additional genes in the sigB operon of Bacillus subtilis that control activity of the general stress factor sigma B in response to environmental signals.
J Bacteriol: 1995, 177(1);123-33
[PubMed:8002610]
[WorldCat.org]
[DOI]
(P p)