Difference between revisions of "GutB"
| Line 15: | Line 15: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU06150 gutB] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU06150 gutB] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/ | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/subtipathways/search.php?enzyme=gutB gutB]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 38 kDa, 5.213 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 38 kDa, 5.213 | ||
Revision as of 10:16, 7 January 2014
- Description: D-sorbitol dehydrogenase
| Gene name | gutB |
| Synonyms | |
| Essential | no |
| Product | D-sorbitol dehydrogenase |
| Function | glucitol utilization |
| Gene expression levels in SubtiExpress: gutB | |
| Metabolic function and regulation of this protein in SubtiPathways: gutB | |
| MW, pI | 38 kDa, 5.213 |
| Gene length, protein length | 1059 bp, 353 aa |
| Immediate neighbours | gutR, gutP |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 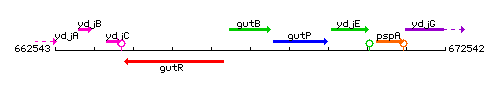
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed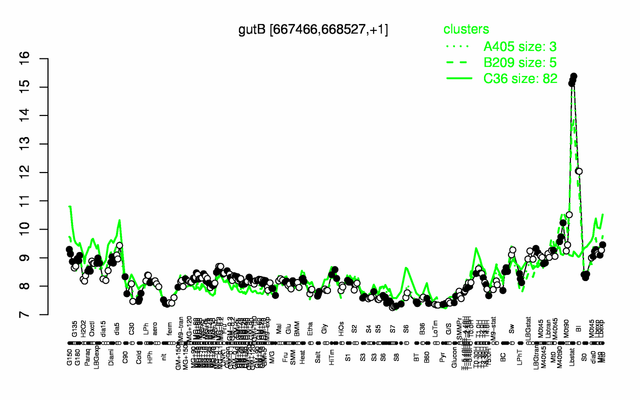
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU06150
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-iditol + NAD+ = L-sorbose + NADH (according to Swiss-Prot)
- Protein family: zinc-containing alcohol dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: Q06004
- KEGG entry: [3]
- E.C. number: 1.1.1.14
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Shouji Watanabe, Miyuki Hamano, Hiroshi Kakeshita, Keigo Bunai, Shigeo Tojo, Hirotake Yamaguchi, Yasutaro Fujita, Sui-Lam Wong, Kunio Yamane
Mannitol-1-phosphate dehydrogenase (MtlD) is required for mannitol and glucitol assimilation in Bacillus subtilis: possible cooperation of mtl and gut operons.
J Bacteriol: 2003, 185(16);4816-24
[PubMed:12897001]
[WorldCat.org]
[DOI]
(P p)
R Ye, S L Wong
Transcriptional regulation of the Bacillus subtilis glucitol dehydrogenase gene.
J Bacteriol: 1994, 176(11);3314-20
[PubMed:8195086]
[WorldCat.org]
[DOI]
(P p)
K Ng, R Ye, X C Wu, S L Wong
Sorbitol dehydrogenase from Bacillus subtilis. Purification, characterization, and gene cloning.
J Biol Chem: 1992, 267(35);24989-94
[PubMed:1460002]
[WorldCat.org]
(P p)