Difference between revisions of "XynA"
| Line 138: | Line 138: | ||
<pubmed>20735481 </pubmed> | <pubmed>20735481 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | + | <pubmed>19531602 ,8012596, 18957862, 19422059 19994888 20096384 20188130 21466172 8628241 20525796 21966517 21785938 24271172 21964501 23459129 </pubmed> | |
| − | <pubmed>19531602 ,8012596, 18957862, 19422059 19994888 20096384 20188130 21466172 8628241 20525796 21966517 21964501 23459129 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:28, 28 November 2013
- Description: endo-1,4-beta-xylanase
| Gene name | xynA |
| Synonyms | |
| Essential | no |
| Product | endo-1,4-beta-xylanase |
| Function | xylan degradation |
| Gene expression levels in SubtiExpress: xynA | |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 23 kDa, 9.644 |
| Gene length, protein length | 639 bp, 213 aa |
| Immediate neighbours | pps, yozV |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 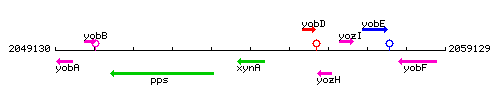
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed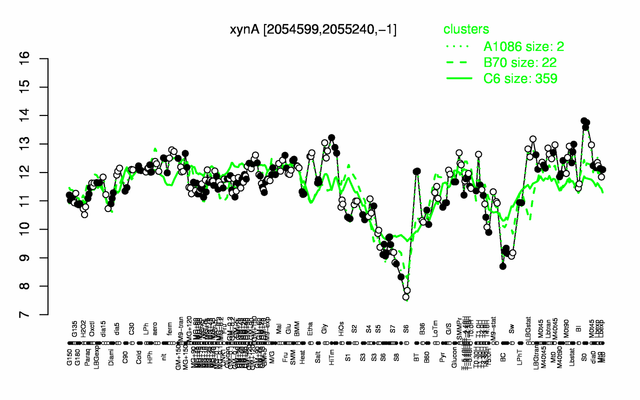
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18840
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endohydrolysis of (1->4)-beta-D-xylosidic linkages in xylans (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- extracellular (signal peptide) PubMed
Database entries
- UniProt: P18429
- KEGG entry: [3]
- E.C. number: 3.2.1.8
Additional information
Expression and regulation
- Operon: xynA PubMed
- Regulation:
- constitutively expressed PubMed
- Regulatory mechanism:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Massimiliano Marvasi, Pieter T Visscher, Lilliam Casillas Martinez
Exopolymeric substances (EPS) from Bacillus subtilis: polymers and genes encoding their synthesis.
FEMS Microbiol Lett: 2010, 313(1);1-9
[PubMed:20735481]
[WorldCat.org]
[DOI]
(I p)
Original publications