Difference between revisions of "Icd"
(→Extended information on the protein) |
|||
| Line 16: | Line 16: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU29130 icd] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU29130 icd] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=Icd Icd] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/carbon_flow.html Central C-metabolism]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/carbon_flow.html Central C-metabolism]''' | ||
Revision as of 09:06, 12 November 2013
- Description: isocitrate dehydrogenase
| Gene name | icd |
| Synonyms | citC |
| Essential | no |
| Product | isocitrate dehydrogenase |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: icd | |
| Interactions involving this protein in SubtInteract: Icd | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 46 kDa, 4.833 |
| Gene length, protein length | 1269 bp, 423 aa |
| Immediate neighbours | mdh, citZ |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 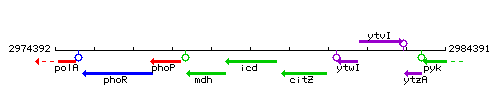
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed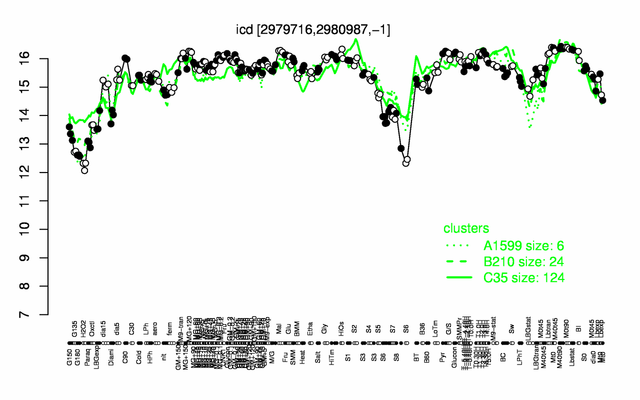
| |
Contents
Categories containing this gene/protein
carbon core metabolism, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29130
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Isocitrate + NADP+ = 2-oxoglutarate + CO2 + NADPH (according to Swiss-Prot)
- Protein family: isocitrate and isopropylmalate dehydrogenases family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: Reversible Michaelis-Menten PubMed
- Domains:
- Modification:
- Cofactor(s): Mg2+, Mn2+, NADP+
- Effectors of protein activity:
- Localization: attached to the membrane PubMed
Database entries
- Structure: 1HQS
- UniProt: P39126
- KEGG entry: [3]
- E.C. number: 1.1.1.42
Additional information
- This enzyme requires NADP+ exclusively. No activity was seen on the presence on NAD+ PubMed
- extensive information on the structure and enzymatic properties of Icd can be found at Proteopedia
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP666 (spc), GP672 (erm), available in Jörg Stülke's lab
- Expression vector:
- pGP1121 (N-terminal Strep-tag, for SPINE, purification from B. subtilis, in pGP380) (available in Jörg Stülke's lab)
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody: available in Linc Sonenshein's lab
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Your additional remarks
References
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Frederik M Meyer, Jan Gerwig, Elke Hammer, Christina Herzberg, Fabian M Commichau, Uwe Völker, Jörg Stülke
Physical interactions between tricarboxylic acid cycle enzymes in Bacillus subtilis: evidence for a metabolon.
Metab Eng: 2011, 13(1);18-27
[PubMed:20933603]
[WorldCat.org]
[DOI]
(I p)
Nico Pietack, Dörte Becher, Sebastian R Schmidl, Milton H Saier, Michael Hecker, Fabian M Commichau, Jörg Stülke
In vitro phosphorylation of key metabolic enzymes from Bacillus subtilis: PrkC phosphorylates enzymes from different branches of basic metabolism.
J Mol Microbiol Biotechnol: 2010, 18(3);129-40
[PubMed:20389117]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Alain Lévine, Françoise Vannier, Cédric Absalon, Lauriane Kuhn, Peter Jackson, Elaine Scrivener, Valérie Labas, Joëlle Vinh, Patrick Courtney, Jérôme Garin, Simone J Séror
Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes.
Proteomics: 2006, 6(7);2157-73
[PubMed:16493705]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Satinder K Singh, Stephen P Miller, Antony Dean, Leonard J Banaszak, David C LaPorte
Bacillus subtilis isocitrate dehydrogenase. A substrate analogue for Escherichia coli isocitrate dehydrogenase kinase/phosphatase.
J Biol Chem: 2002, 277(9);7567-73
[PubMed:11751849]
[WorldCat.org]
[DOI]
(P p)
S K Singh, K Matsuno, D C LaPorte, L J Banaszak
Crystal structure of Bacillus subtilis isocitrate dehydrogenase at 1.55 A. Insights into the nature of substrate specificity exhibited by Escherichia coli isocitrate dehydrogenase kinase/phosphatase.
J Biol Chem: 2001, 276(28);26154-63
[PubMed:11290745]
[WorldCat.org]
[DOI]
(P p)
C Jourlin-Castelli, N Mani, M M Nakano, A L Sonenshein
CcpC, a novel regulator of the LysR family required for glucose repression of the citB gene in Bacillus subtilis.
J Mol Biol: 2000, 295(4);865-78
[PubMed:10656796]
[WorldCat.org]
[DOI]
(P p)
K Matsuno, T Blais, A W Serio, T Conway, T M Henkin, A L Sonenshein
Metabolic imbalance and sporulation in an isocitrate dehydrogenase mutant of Bacillus subtilis.
J Bacteriol: 1999, 181(11);3382-91
[PubMed:10348849]
[WorldCat.org]
[DOI]
(P p)
S Jin, P A Levin, K Matsuno, A D Grossman, A L Sonenshein
Deletion of the Bacillus subtilis isocitrate dehydrogenase gene causes a block at stage I of sporulation.
J Bacteriol: 1997, 179(15);4725-32
[PubMed:9244258]
[WorldCat.org]
[DOI]
(P p)
R F Ramaley, M O Hudock
Purification and properties of isocitrate dehydrogenase (NADP) from Thermus aquaticus YT-1, Bacillus subtilis-168 and Chlamydomonas reinhardti-Y-2.
Biochim Biophys Acta: 1973, 315(1);22-36
[PubMed:4147570]
[WorldCat.org]
[DOI]
(P p)