Difference between revisions of "RnhC"
(→Biological materials) |
|||
| Line 115: | Line 115: | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' BP430 (''ΔrnhC::cat'') available in |
* '''Expression vector:''' | * '''Expression vector:''' | ||
Revision as of 15:35, 29 October 2013
- Description: RNase HIII, endoribonuclease
| Gene name | rnhC |
| Synonyms | ysgB |
| Essential | no |
| Product | Mg2+-dependent RNase HIII |
| Function | endonucleolytic cleavage of RNA in RNA-DNA hybrid molecules |
| Gene expression levels in SubtiExpress: rnhC | |
| MW, pI | 33 kDa, 10.07 |
| Gene length, protein length | 939 bp, 313 aa |
| Immediate neighbours | zapA, pheT |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 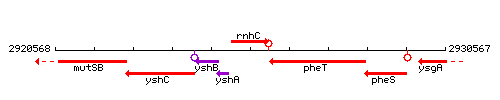
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed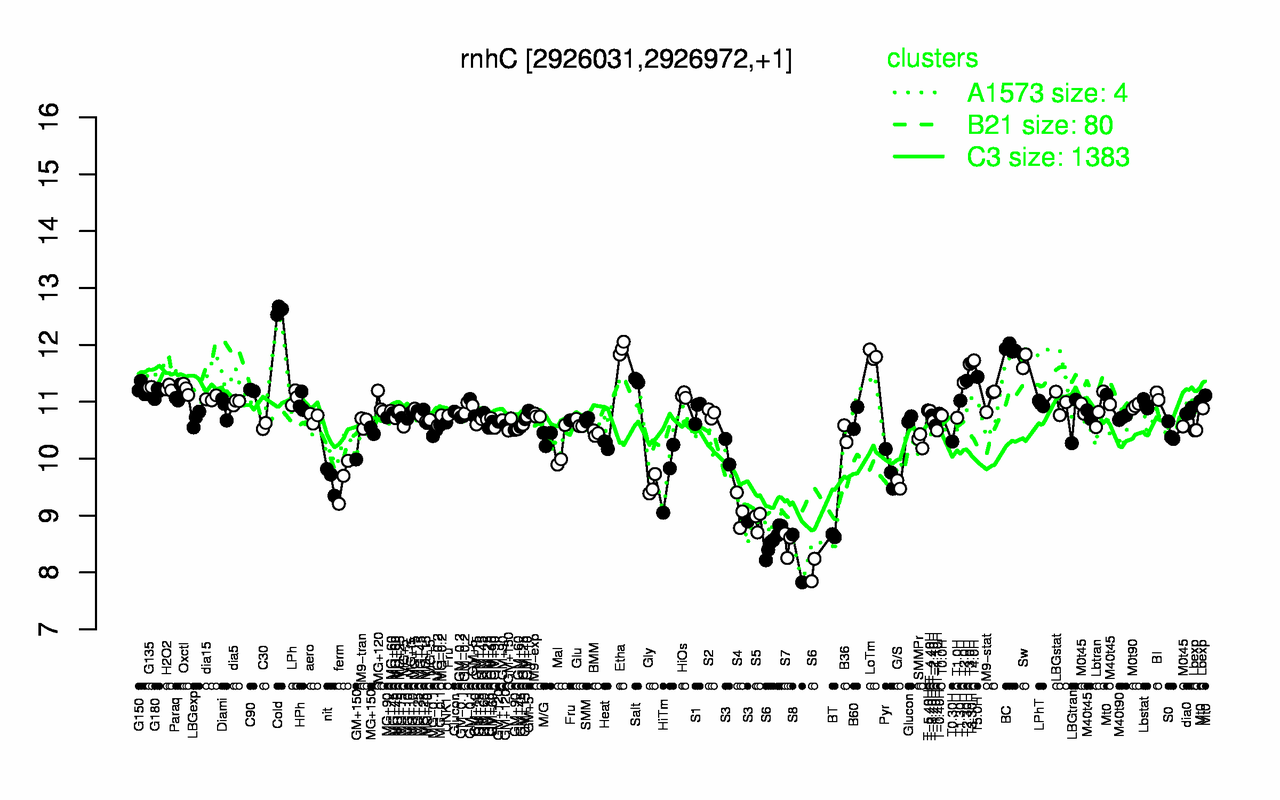
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28620
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Endonucleolytic cleavage to 5'-phosphomonoester (according to Swiss-Prot)
- Protein family: RnhC subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- RnhC interacts with the RNA polymerase PubMed
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 2D0B (complex with Mg2+, Geobacillus stearothermophilus , 47% identity), 2D0A (Geobacillus stearothermophilus, 47% identity)
- UniProt: P94541
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: BP430 (ΔrnhC::cat) available in
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications